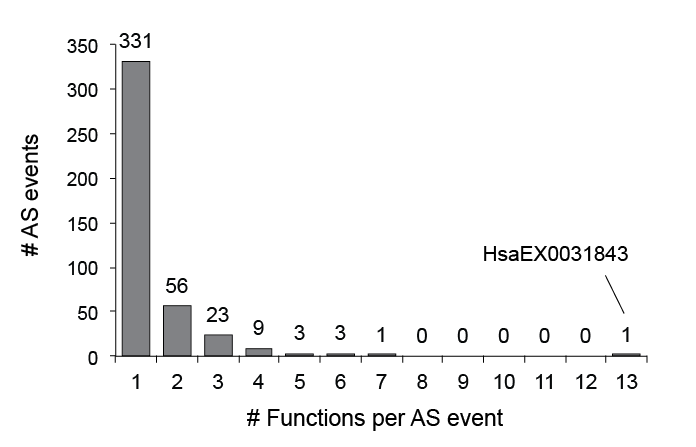
Functions v1.2 byevent.png
From VastDB
Revision as of 13:14, 26 November 2020 by WikiSysop (talk | contribs) (WikiSysop uploaded File:Functions v1.2 byevent.png)
Functions_v1.2_byevent.png (696 × 447 pixels, file size: 17 KB, MIME type: image/png)
File history
Click on a date/time to view the file as it appeared at that time.
| Date/Time | Thumbnail | Dimensions | User | Comment | |
|---|---|---|---|---|---|
| current | 15:12, 3 September 2018 |  | 696 × 447 (17 KB) | WikiSysop (talk | contribs) |
- You cannot overwrite this file.
File usage
The following page links to this file: