GgaEX6014719 @ galGal3
Exon Skipping
Gene
ENSGALG00000008984 | C9orf86
Description
NA
Coordinates
chr17:965399-968100:-
Coord C1 exon
chr17:967929-968100
Coord A exon
chr17:966096-966145
Coord C2 exon
chr17:965399-965476
Length
50 bp
Sequences
Splice sites
3' ss Seq
CTGTTTGTTTGTTTGAACAGATA
3' ss Score
9.09
5' ss Seq
GAGGTATTA
5' ss Score
6.46
Exon sequences
Seq C1 exon
GATGAATTCCCTGTCCGGGAAGATCTCTCTGAAATATCTGATGATGATACCTCATTGGCAAAGCCACCGCAGCCTGTGAAATCAACAGTCCACTCCTTCAAACTGAAAAATGATTCAGACCTCTTTGGTTTGGGTTTGGAAGAAACCGGTCCCAAGGAGAGCAGCGAAGAAG
Seq A exon
ATAAACACGCTTCTAAGGAGAAGAAGAAGAAAAAGAAGAAAAGCAAAGAG
Seq C2 exon
GAAGAGGAGAAGTCTGTGAAAAAGAAGAGCAAACACAAGAAGGGCAAAGAGAAAGAGGAGAGCAAAGAGGAGAAAGAC
VastDB Features
Vast-tools module Information
Secondary ID
ENSGALG00000008984-'24-28,'24-27,25-28=AN
Average complexity
A_S
Mappability confidence:
100%=100=100%
Protein Impact
NA
No structure available
Features
Disorder rate (Iupred):
C1=0.983 A=1.000 C2=1.000
Domain overlap (PFAM):
C1:
NO
A:
NO
C2:
NO
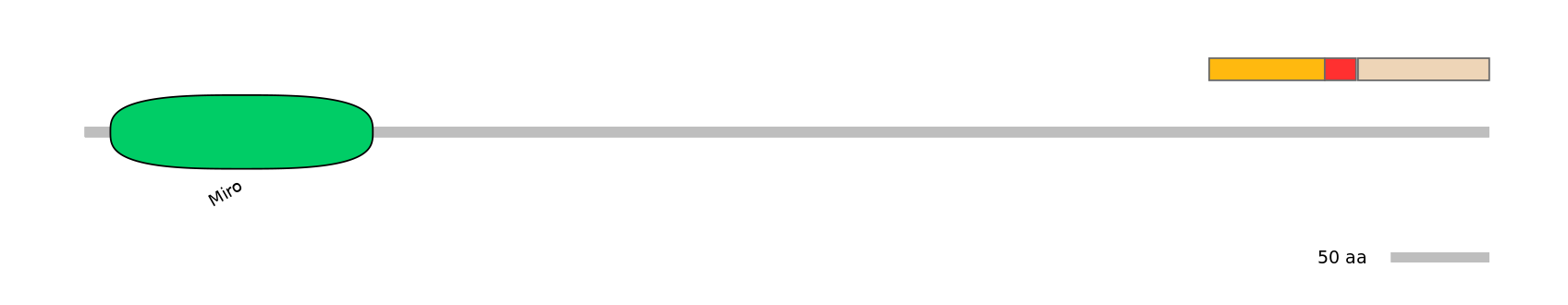
Main Skipping Isoform:
NA
Other Inclusion Isoforms:
NA
Other Skipping Isoforms:
NA
Associated events
Other assemblies
Conservation
Human
(hg19)
No conservation detected
Mouse
(mm9)
No conservation detected
Fruitfly
(dm6)
No conservation detected
Primers PCR
Suggestions for RT-PCR validation
F:
AGCCTGTGAAATCAACAGTCCA
R:
TGCCCTTCTTGTGTTTGCTCT
Band lengths:
148-198
Functional annotations
There are 1 annotated functions for this event
PMID: 19433581
All four RBEL1 isoforms (A, B, C, and D) have identical N termini harboring the Rab-like GTPase domains but contain variable C termini. Although all isoforms can be detected in both cytoplasm and nucleus, RBEL1A is predominantly cytoplasmic, whereas RBEL1B is mostly nuclear. RBEL1C and -D, by contrast, are evenly distributed between the cytoplasm and nucleus. Furthermore, all four RBEL1 proteins are also capable of associating with cellular membrane. The RBEL1 proteins also exhibit a unique nucleotide-binding potential and, whereas the larger A and B isoforms are mainly GTP-bound, the smaller C and D variants bind to both GTP and GDP. Furthermore, a regulatory region at amino acid position 236-302 immediately adjacent to the GTP-binding domain is important for GTP-binding potential of RBEL1A, because deletion of this region converts RBEL1A from predominantly GTP-bound to GDP-bound. RBEL1 knockdown via RNA interference results in marked cell growth suppression, which is associated with morphological and biochemical features of apoptosis as well as inhibition of extracellular signal-regulated kinase phosphorylation. Notes: C and D: internal APA, the difference is HsaINT0025589 (retained in D). B: ALTD (ex 9, HsaALTD0001096) top ALTA (ex 15, not in VastDB) skipping exons 10 (not in VastDB), 11 (HsaEX1030457), 12 (HsaEX7005708), 13 (HsaEX6064086), 14 (HsaEX6064085).
GENOMIC CONTEXT[edit]
INCLUSION PATTERN[edit]