HsaALTD0007408-2/2 @ hg38
Alternative 5'ss
Gene
ENSG00000137871 | ZNF280D
Description
zinc finger protein 280D [Source:HGNC Symbol;Acc:HGNC:25953]
Coordinates
chr15:56666395-56666986:-
Coord C1 exon
chr15:56666800-56666986
Coord A exon
chr15:56666679-56666799
Coord C2 exon
chr15:56666395-56666535
Length
121 bp
Sequences
Splice sites
5' ss Seq
AAGGTAATC
5' ss Score
7.46
3' ss Seq
TTTTTTAATGTCTCCCTAAGGTA
3' ss Score
7.73
Exon sequences
Seq C1 exon
GTTACTATTCGAGCTTCAGTTGGACCTCTGCAATCAGGAGCTTCACCTACACCTTCCATTAGTGCAAGTGCTTCTACCCTTCAGCTCTCACCTCCAAGGACTAAAAATATAACTGCTAAAAATCCTGCAAAATCTAATACAAGTAAACCTAATACAGTCAAATCCAACGCAAGTAAACCTAATACAA
Seq A exon
GTAAGCCTAATGGAAGTAAATCTAAATACAAACCAAAAATCTCTAATATGCAAAAGAAACAAAGCACATTGGCTAGTAGCAATAAAAAAAGTAAAGTCAATACAGCATTGAGGAATTTAAG
Seq C2 exon
GTATCGTCGGGGCATTCACAAATGCATTGAGTGTTGTTCCGAAATAAAAGATTTTGCAAACCACTTTCCTACGTACGTCCACTGTAGTTTTTGCAGATACAACACTAGCTGTAGCAAAGCCTATGTAAATCATATGATGAG
VastDB Features
Vast-tools module Information
Secondary ID
ENSG00000137871-55-71,56-71-2/2
Average complexity
Alt5
Mappability confidence:
NA
Protein Impact
Protein isoform when splice site is used (Ref)
No structure available
Features
Disorder rate (Iupred):
C1=0.957 A=NA C2=0.013
Domain overlap (PFAM):
C1:
NO
A:
NA
C2:
NO
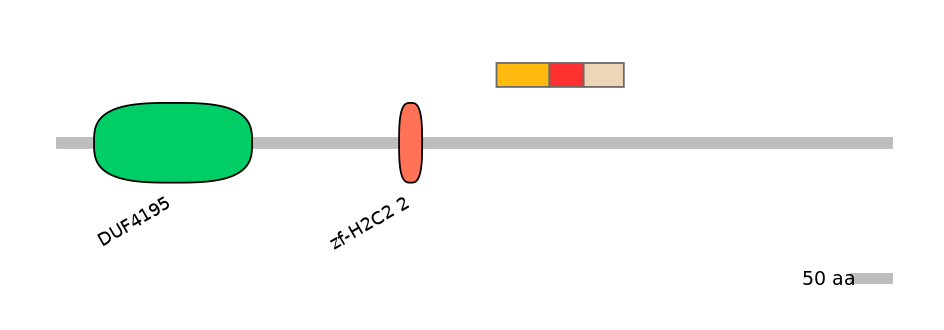
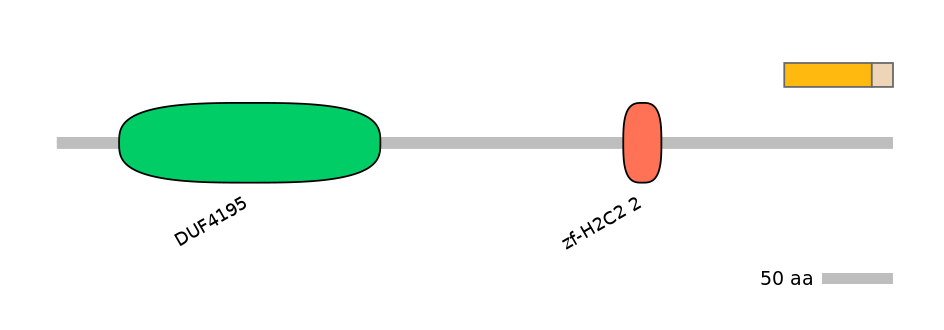
Other Skipping Isoforms:
NA
Associated events
Other assemblies
Conservation
Mouse
(mm9)
No conservation detected
Cow
(bosTau6)
No conservation detected
Chicken
(galGal3)
No conservation detected
Fruitfly
(dm6)
No conservation detected
Primers PCR
Suggestions for RT-PCR validation
F:
GAGCTTCAGTTGGACCTCTGC
R:
TGGACGTACGTAGGAAAGTGGT
Band lengths:
258-379
Functional annotations
There are 0 annotated functions for this event
GENOMIC CONTEXT[edit]
INCLUSION PATTERN[edit]
SPECIAL DATASETS
- The Cancer Genome Atlas (TCGA)
- Autistic and control brains
- Pre-implantation embryo development