HsaALTD1020895-2/2 @ hg38
Alternative 5'ss
Gene
ENSG00000135407 | AVIL
Description
advillin [Source:HGNC Symbol;Acc:HGNC:14188]
Coordinates
chr12:57808395-57809695:-
Coord C1 exon
chr12:57809597-57809695
Coord A exon
chr12:57809542-57809596
Coord C2 exon
chr12:57808395-57808548
Length
55 bp
Sequences
Splice sites
5' ss Seq
CAGGTGTGT
5' ss Score
6.99
3' ss Seq
GGGAATGGGTGGCTCTGCAGGGC
3' ss Score
3.48
Exon sequences
Seq C1 exon
GACTGCTACATCCTGGACCAAAGTGGAACCAAAATCTACGTGTGGAAAGGAAAAGGAGCCACAAAGGCTGAAAAACAGGCAGCCATGTCTAAAGCGCTG
Seq A exon
GTAGGAGCTACAGATGTGCAATACTGTTCAAATAATTCTGCTAAGAGCACTTCAG
Seq C2 exon
GGCTTCATCAAGATGAAGAGCTACCCCAGCAGCACCAATGTGGAGACCGTCAACGATGGTGCTGAGTCGGCCATGTTCAAGCAGCTGTTCCAGAAGTGGTCAGTAAAGGACCAGACCATGGGCCTGGGGAAAACGTTCAGCATTGGTAAAATTG
VastDB Features
Vast-tools module Information
Secondary ID
ENSG00000135407-17-17,18-17-2/2
Average complexity
Alt5
Mappability confidence:
NA
Protein Impact
ORF disruption when splice site is used (sequence inclusion)
No structure available
Features
Disorder rate (Iupred):
C1=0.000 A=NA C2=0.019
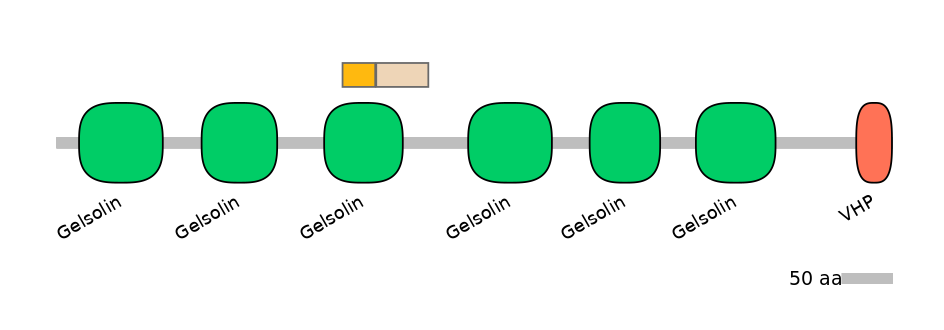
Domain overlap (PFAM):
C1:
PF0062617=Gelsolin=FE(41.0=100)
A:
NA
C2:
PF0062617=Gelsolin=PD(33.3=50.0)
Main Inclusion Isoform:
NA
Other Inclusion Isoforms:
NA
Other Skipping Isoforms:
NA
Associated events
Conservation
Mouse
(mm9)
No conservation detected
Cow
(bosTau6)
No conservation detected
Chicken
(galGal4)
No conservation detected
Chicken
(galGal3)
No conservation detected
Zebrafish
(danRer10)
No conservation detected
Fruitfly
(dm6)
No conservation detected
Primers PCR
Suggestions for RT-PCR validation
F:
ACAGGCAGCCATGTCTAAAGC
R:
GCCCATGGTCTGGTCCTTTAC
Band lengths:
148-203
Functional annotations
There are 0 annotated functions for this event
GENOMIC CONTEXT[edit]
INCLUSION PATTERN[edit]
SPECIAL DATASETS
- Autistic and control brains