HsaALTD0001450-4/5 @ hg38
Alternative 5'ss
Gene
ENSG00000148337 | CIZ1
Description
CDKN1A interacting zinc finger protein 1 [Source:HGNC Symbol;Acc:HGNC:16744]
Coordinates
chr9:128178709-128179415:-
Coord C1 exon
chr9:128179373-128179415
Coord A exon
chr9:128179085-128179372
Coord C2 exon
chr9:128178709-128178904
Length
288 bp
Sequences
Splice sites
5' ss Seq
CAGGTGCAG
5' ss Score
4.95
3' ss Seq
AGACACAGACATATCCACAGGTC
3' ss Score
1.02
Exon sequences
Seq C1 exon
CTCAGAAGAGCCCACAGAGAAGGAACCTCCAGGGCAGTTACAG
Seq A exon
GTGAAGGCCCAGCCGCAGGCCCGGATGACAGTACCGAAACAGACACAGACACCAGACCTGCTGCCTGAGGCCCTGGAAGCCCAAGTGCTGCCACGATTCCAGCCACGGGTCCTGCAGGTCCAGGCCCAGGTGCAGTCACAGACTCAGCCGCGGATACCATCCACAGACACCCAGGTGCAGCCAAAGCTGCAGAAGCAGGCGCAAACACAGACCTCTCCAGAGCACTTAGTGCTGCAACAGAAGCAGGTGCAGCCACAGCTGCAGCAGGAGGCAGAGCCACAGAAGCAG
Seq C2 exon
GTCCACACACAGGCACAGCCAAGCGTCCAGCCACAGGAGCATCCTCCAGCGCAGGTGTCAGTACAGCCACCAGAGCAGACCCATGAGCAGCCTCACACCCAGCCGCAGGTGTCGTTGCTGGCTCCAGAGCAAACACCAGTTGTGGTTCATGTCTGCGGGCTGGAGATGCCACCTGATGCAGTAGAAGCTGGTGGAG
VastDB Features
Vast-tools module Information
Secondary ID
ENSG00000148337-35-75,36-75,37-75,41-75,42-75-4/5
Average complexity
Alt5
Mappability confidence:
NA
Protein Impact
Protein isoform when splice site is used (No Ref)
No structure available
Features
Disorder rate (Iupred):
C1=1.000 A=NA C2=0.967
Domain overlap (PFAM):
C1:
NO
A:
NA
C2:
NO
Main Inclusion Isoform:
NA
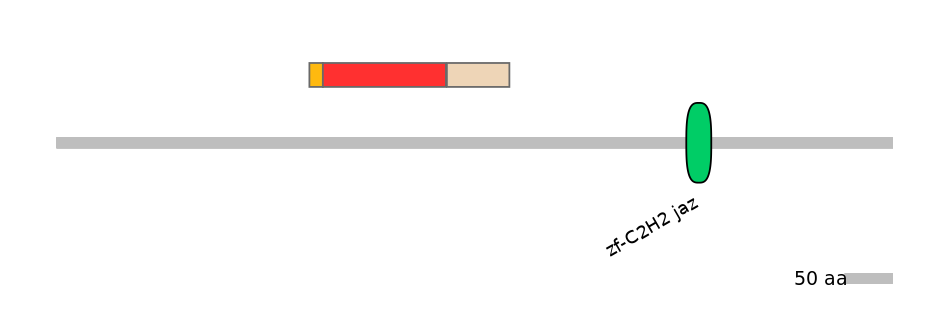
Other Inclusion Isoforms:
NA
Associated events
Conservation
Mouse
(mm10)
No conservation detected
Mouse
(mm9)
No conservation detected
Cow
(bosTau6)
No conservation detected
Chicken
(galGal3)
No conservation detected
Fruitfly
(dm6)
No conservation detected
Primers PCR
Suggestions for RT-PCR validation
F:
AGAAGAGCCCACAGAGAAGGA
R:
CTCCACCAGCTTCTACTGCAT
Band lengths:
236-524
Functional annotations
There are 0 annotated functions for this event
GENOMIC CONTEXT[edit]
INCLUSION PATTERN[edit]
SPECIAL DATASETS
- Autistic and control brains