HsaEX0053044 @ hg38
Exon Skipping
Gene
ENSG00000214022 | REPIN1
Description
replication initiator 1 [Source:HGNC Symbol;Acc:HGNC:17922]
Coordinates
chr7:150369671-150374039:+
Coord C1 exon
chr7:150369671-150369868
Coord A exon
chr7:150370760-150370884
Coord C2 exon
chr7:150371228-150374039
Length
125 bp
Sequences
Splice sites
3' ss Seq
TTGCTTCATCTGTTAAACAGGGA
3' ss Score
7.22
5' ss Seq
CTGGTGAGT
5' ss Score
10.1
Exon sequences
Seq C1 exon
GTAAGGCTGGCCTCTCTGCAGTCAGAGGTCTGAGCTCTGCCATGGGGATAGGGGTGTCTTTATTACTGCAGTTTTCTCTAACACCTGGGGGCTACCGGAGTGTGGGCCGAAGCAGGCGCTGCAGCCGCGGAAGTATCCCCAGGAACATCCCCAAGAGGAGCTGGAAAAAGCCTCATCCCCAGCTCTGCAGTCTCCAGG
Seq A exon
GGAGCTCAGTGTCTGTTTGTCCAGCTTCTCAGAGTTGCTGTGCAGCTCGGATGTGGCATAGGAAACAGCAGACACAGGGAGAGGGCAGCATAAGGCACTGTAGGGAGCAGTGGCCACATTTTCTG
Seq C2 exon
CAGAGGAAGAACCGATGCTGGAACGTCGTTGCAGGGGCCCCCTGGCCATGGGCCTGGCCCAGCCCCGACTCCTTTCTGGGCCCTCCCAGGAGTCACCCCAGACCCTGGGGAAGGAGTCCCGCGGGCTGAGGCAACAAGGCACGTCAGTGGCCCAGTCTGGTGCCCAAGCCCCAGGCAGGGCCCATCGCTGTGCCCACTGTCGAAGGCACTTCCCTGGCTGGGTGGCTCTGTGGCTTCACACCCGCCGGTGCCAGGCCCGGCTGCCCTTGCCCTGCCCTGAGTGTGGCCGTCGCTTTCGCCATGCCCCCTTCTTAGCACTGCACCGCCAGGTCCATGCTGCTGCCACCCCAGACCTGGGCTTTGCCTGCCACCTCTGTGGGCAGAGCTTCCGAGGCTGGGTGGCCCTGGTTCTGCATCTGCGGGCCCATTCAGCTGCAAAGCGGCCCATCGCTTGTCCCAAATGCGAGAGACGCTTCTGGCGACGAAAGCAGCTTCGAGCT
VastDB Features
Vast-tools module Information
Secondary ID
ENSG00000214022_MULTIEX1-2/2=1-C2
Average complexity
S*
Mappability confidence:
100%=100=100%
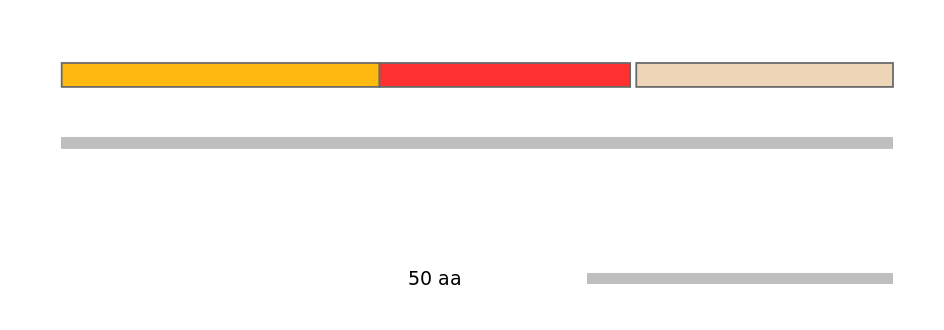
Protein Impact
Alternative protein isoforms (No Ref, Alt. Stop)
No structure available
Features
Disorder rate (Iupred):
C1=0.355 A=0.125 C2=0.401
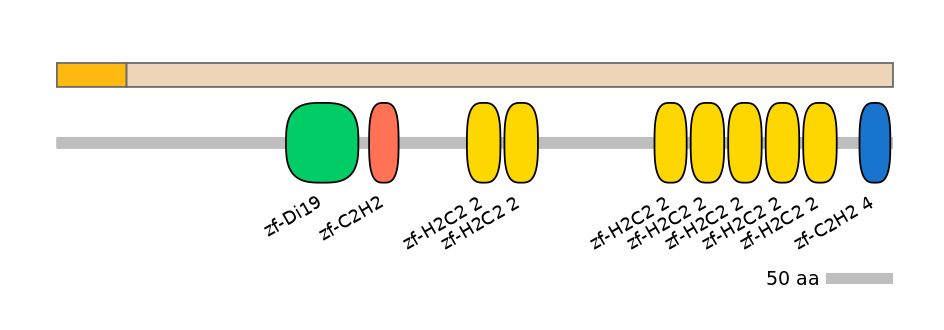
Domain overlap (PFAM):
C1:
NO
A:
NO
C2:
PF0009621=zf-C2H2=WD(100=4.0),PF138941=zf-C2H2_4=WD(100=4.2),PF0009621=zf-C2H2=WD(100=4.0),PF134651=zf-H2C2_2=WD(100=4.6),PF134651=zf-H2C2_2=WD(100=4.6),PF134651=zf-H2C2_2=WD(100=4.4),PF134651=zf-H2C2_2=WD(100=4.6),PF134651=zf-H2C2_2=WD(100=4.6),PF134651=zf-H2C2_2=WD(100=4.6),PF134651=zf-H2C2_2=WD(100=4.6),PF138941=zf-C2H2_4=WD(100=4.2)
Associated events
Other assemblies
Conservation
Mouse
(mm9)
No conservation detected
Cow
(bosTau6)
No conservation detected
Chicken
(galGal3)
No conservation detected
Zebrafish
(danRer10)
No conservation detected
Fruitfly
(dm6)
No conservation detected
Primers PCR
Suggestions for RT-PCR validation
F:
AAAAAGCCTCATCCCCAGCTC
R:
GAAGTGCCTTCGACAGTGGG
Band lengths:
246-371
Functional annotations
There are 0 annotated functions for this event
GENOMIC CONTEXT[edit]
INCLUSION PATTERN[edit]
SPECIAL DATASETS
- The Cancer Genome Atlas (TCGA)
- Genotype-Tissue Expression Project (GTEx)
- Autistic and control brains
- Pre-implantation embryo development