HsaEX0066779 @ hg19
Exon Skipping
Gene
ENSG00000164548 | TRA2A
Description
transformer 2 alpha homolog (Drosophila) [Source:HGNC Symbol;Acc:16645]
Coordinates
chr7:23561326-23571660:-
Coord C1 exon
chr7:23571408-23571660
Coord A exon
chr7:23561740-23562051
Coord C2 exon
chr7:23561326-23561459
Length
312 bp
Sequences
Splice sites
3' ss Seq
AATGTTCTTTTTCCATTAAGGTT
3' ss Score
8.88
5' ss Seq
AATGTATGG
5' ss Score
3.37
Exon sequences
Seq C1 exon
GCCCGCCCCCTTTACGTCGTGCTCTGAGGAGGCCAGGAGATTTCTGGCGGCGCCGGCGCCATTTTGCTGGAGCCTGCGACCGAGTGGGAGTGGAGTGGAGCGGCTGTGGTTGCCGACTCTTTCCTCTTCCCCACGGTCCAGTCAGCGGGTTAATTAGGCCATCGGCCCTCGAGCCGAGACTTGTCTCTTATTTAGTTCTGGGGAGCGCCTCGTCGACATGAGTGATGTGGAGGAAAACAACTTCGAGGGCAGA
Seq A exon
GTTAATGTTCGTGAAGAAATTGAAGAGTTTTTTCCAAGAATGTGGAAGATAAATCAAGATAAAAGAAGGCTAATGAAAAGTATTAAAGATCAGAAAATTAAAATTGAATGGGGGAAAAAATTGAACAGAAGATTGGTCAGATAGAAGCACTTGAATATTTTTTTAAGGCTATTTTGAATTGTTTGAATTGGGGAAGAATACACGAAGTATGAAAAATGAAAAACTCAATGAAGAATGAAGAGAAGTAGAAAGCAAGAGTGAAGTAGAATTAAAAGAATCTGGAAGAATGAATGGGTCCTAGGTTAAGTAAAT
Seq C2 exon
GAGTCTCGCTCTCAGTCAAAATCTCCAACGGGAACTCCTGCTCGTGTAAAATCGGAGAGCAGGTCAGGATCTCGTAGTCCATCAAGGGTTTCCAAACACTCTGAATCCCATTCTCGATCAAGATCAAAATCCAG
VastDB Features
Vast-tools module Information
Secondary ID
ENSG00000164548_MULTIEX1-2/3=C1-3
Average complexity
C1*
Mappability confidence:
100%=100=100%
Protein Impact
In the CDS, with uncertain impact
No structure available
Features
Disorder rate (Iupred):
C1=0.429 A=0.000 C2=0.643
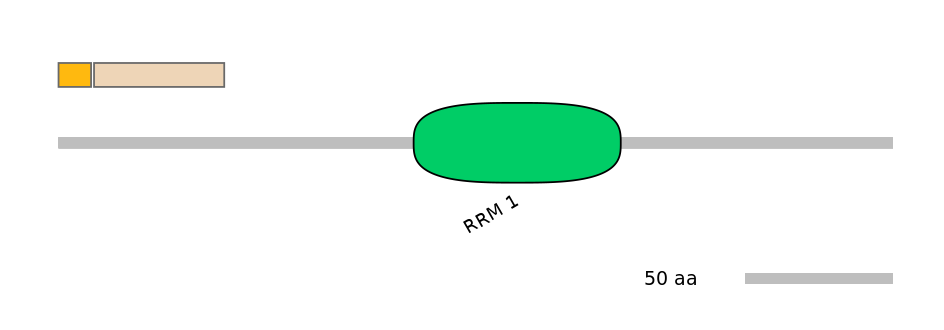
Domain overlap (PFAM):
C1:
NO
A:
NO
C2:
NO
Other Inclusion Isoforms:
NA
Other Skipping Isoforms:
NA
Associated events
Other assemblies
Conservation
Fruitfly
(dm6)
No conservation detected
Primers PCR
Suggestions for RT-PCR validation
F:
CCTTTACGTCGTGCTCTGAGG
R:
TGTTTGGAAACCCTTGATGGACT
Band lengths:
343-655
Functional annotations
There are 3 annotated functions for this event
PMID: 30916337
This event
Experimentally validated NMD (response to KD of NMD machinery; from 18443041) and eCLIP binding in an autoregulatory loop.
PMID: 31911676
This event
[CRISPR screen]. Conserved poison exon with mixed impact when depleted: HeLa Enriched at 14 days (FC=1.256, FDR=0.002), PC9 No Change at 14 days (FC=1.256, FDR=0.158), Late Xenograft Depleted (FC=0.604, FDR=0.000).
PMID: 33176162
This event
Poison exons in serine-arginine-rich (SR) proteins, a family of 14 essential SFs, are differentially spliced during induced pluripotent stem cell (iPSC) differentiation and in tumors versus normal tissues. The study uncovers an extensive cross-regulatory network of SR proteins controlling their expression via alternative splicing coupled to nonsense-mediated decay. This TRA2A PE is in the CDS.
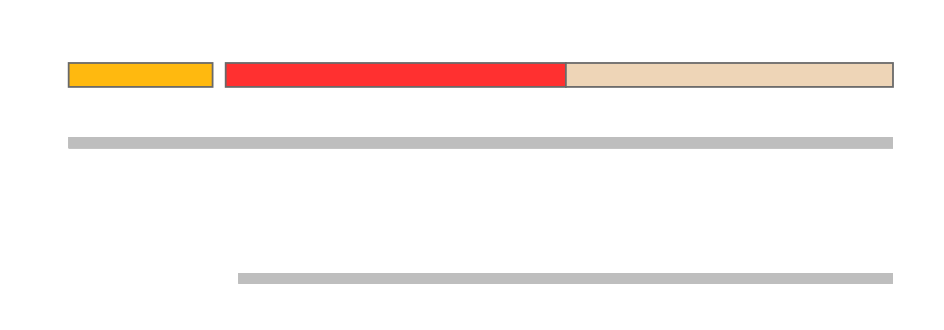
GENOMIC CONTEXT[edit]
INCLUSION PATTERN[edit]
SPECIAL DATASETS
- The Cancer Genome Atlas (TCGA)
- Genotype-Tissue Expression Project (GTEx)
- Autistic and control brains
- Pre-implantation embryo development
Other AS DBs:
FasterDB (Includes CLIP-seq data)
AS-ALPS (AS-induced ALteration of Protein Structure, links to PINs)
APPRIS (Selection of principal isoform)
DEU primates (Only for human)