HsaEX0070745 @ hg19
Exon Skipping
Gene
ENSG00000165816 | VWA2
Description
von Willebrand factor A domain containing 2 [Source:HGNC Symbol;Acc:24709]
Coordinates
chr10:116045698-116049751:+
Coord C1 exon
chr10:116045698-116046270
Coord A exon
chr10:116048109-116048290
Coord C2 exon
chr10:116048697-116049751
Length
182 bp
Sequences
Splice sites
3' ss Seq
TTTCATGTCTGTTCCCCTAGACT
3' ss Score
9.53
5' ss Seq
GAGGTGAGT
5' ss Score
10.03
Exon sequences
Seq C1 exon
CGGCTTCCTGCGGGCCAAAGTCTTCGTGAAGCGGTTTGTGCGGGCCGTGCTGAGCGAGGACTCTCGGGCCCGAGTGGGTGTGGCCACATACAGCAGGGAGCTGCTGGTGGCGGTGCCTGTGGGGGAGTACCAGGATGTGCCTGACCTGGTCTGGAGCCTCGATGGCATTCCCTTCCGTGGTGGCCCCACCCTGACGGGCAGTGCCTTGCGGCAGGCGGCAGAGCGTGGCTTCGGGAGCGCCACCAGGACAGGCCAGGACCGGCCACGTAGAGTGGTGGTTTTGCTCACTGAGTCACACTCCGAGGATGAGGTTGCGGGCCCAGCGCGTCACGCAAGGGCGCGAGAGCTGCTCCTGCTGGGTGTAGGCAGTGAGGCCGTGCGGGCAGAGCTGGAGGAGATCACAGGCAGCCCAAAGCATGTGATGGTCTACTCGGATCCTCAGGATCTGTTCAACCAAATCCCTGAGCTGCAGGGGAAGCTGTGCAGCCGGCAGCGGCCAG
Seq A exon
ACTGTCAGCCTGTTGACAGCAGGCATGGGCCCATTTTGTCCATACAGCATCTAATTAGTGCCCTGCATACTGGGGATGGATGAAGGCACAGGTGAAGATGTTGTCACAAAGGGTTGCCCACAAACCCAAACCCCATGGGTTACATAAGACAGGAACTTATTTCTCTCACATACAGTTCAGAG
Seq C2 exon
GGTGCCGGACACAAGCCCTGGACCTCGTCTTCATGTTGGACACCTCTGCCTCAGTAGGGCCCGAGAATTTTGCTCAGATGCAGAGCTTTGTGAGAAGCTGTGCCCTCCAGTTTGAGGTGAACCCTGACGTGACACAGGTCGGCCTGGTGGTGTATGGCAGCCAGGTGCAGACTGCCTTCGGGCTGGACACCAAACCCACCCGGGCTGCGATGCTGCGGGCCATTAGCCAGGCCCCCTACCTAGGTGGGGTGGGCTCAGCCGGCACCGCCCTGCTGCACATCTATGACAAAGTGATGACCGTCCAGAGGGGTGCCCGGCCTGGTGTCCCCAAAGCTGTGGTGGTGCTCACAGGCGGGAGAGGCGCAGAGGATGCAGCCGTTCCTGCCCAGAAGCTGAGGAACAATGGCATCTCTGTCTTGGTCGTGGGCGTGGGGCCTGTCCTAAGTGAGGGTCTGCGGAGGCTTGCAGGTCCCCGGGATTCCCTGATCCACGTGGCAGCT
VastDB Features
Vast-tools module Information
Secondary ID
ENSG00000165816_CASSETTE1
Average complexity
S
Mappability confidence:
100%=100=100%
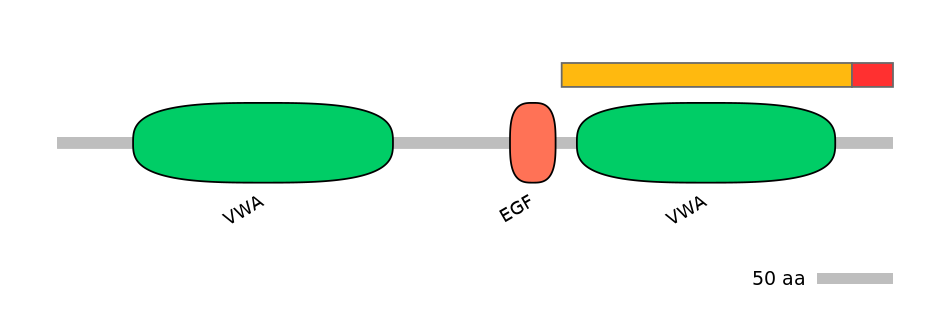
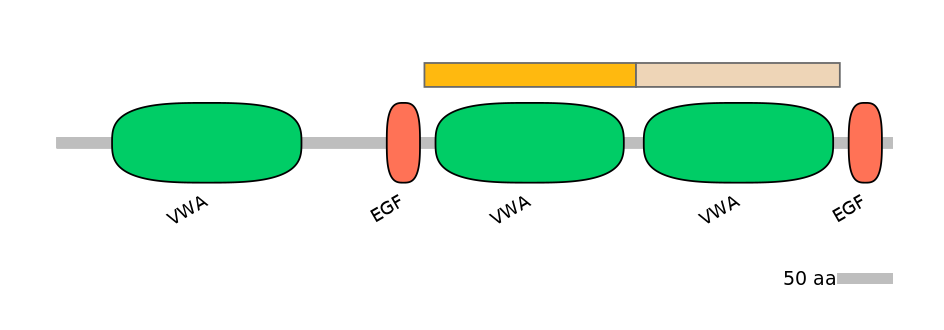
Protein Impact
ORF disruption upon sequence inclusion
No structure available
Features
Disorder rate (Iupred):
C1=0.122 A=0.333 C2=0.061
Domain overlap (PFAM):
C1:
PF0009223=VWA=WD(100=89.1)
A:
NO
C2:
PF0009223=VWA=WD(100=84.7)
Other Inclusion Isoforms:
NA
Associated events
Other assemblies
Conservation
Rat
(rn6)
No conservation detected
Chicken
(galGal4)
No conservation detected
Chicken
(galGal3)
No conservation detected
Zebrafish
(danRer10)
No conservation detected
Fruitfly
(dm6)
No conservation detected
Primers PCR
Suggestions for RT-PCR validation
F:
GAGATCACAGGCAGCCCAAAG
R:
CTGTGTCACGTCAGGGTTCAC
Band lengths:
243-425
Functional annotations
There are 0 annotated functions for this event
GENOMIC CONTEXT[edit]
INCLUSION PATTERN[edit]
SPECIAL DATASETS
- The Cancer Genome Atlas (TCGA)
- Genotype-Tissue Expression Project (GTEx)
- Autistic and control brains
- Pre-implantation embryo development
Other AS DBs:
FasterDB (Includes CLIP-seq data)
AS-ALPS (AS-induced ALteration of Protein Structure, links to PINs)
APPRIS (Selection of principal isoform)
DEU primates (Only for human)