GgaALTA1017171-1/3 @ galGal4
Alternative 3'ss
Gene
ENSGALG00000002451 | KCNT2
Description
potassium channel, sodium activated subfamily T, member 2 [Source:HGNC Symbol;Acc:HGNC:18866]
Coordinates
chr8:2767066-2772717:+
Coord C1 exon
chr8:2767066-2767238
Coord A exon
NA
Coord C2 exon
chr8:2772621-2772717
Length
0 bp
Sequences
Splice sites
5' ss Seq
CCCGTATGT
5' ss Score
5.55
3' ss Seq
ATGTGGGGTCACCTTTGCAGCTA
3' ss Score
6.87
Exon sequences
Seq C1 exon
GGGTGCCAGCATAACTACTATGAAGATGCCAAAGCCTATGGATTCAAAAATAAATTGATTATTGTTGCAGCAGAAACTGCTGGAAATGGCTTATATAATTTTATTGTTCCACTTAGAGCTTATTACAGACCGAAGAAAGAACTCAATCCCATTGTATTACTCCTGGATAACCC
Seq A exon
NA
Seq C2 exon
CTAATATGGTGGTGGTGGACAAAGAAAGTACAATGAGTGCTGAGGAAGACTACATGGCAGATGCTAAGACTATTGTAAACGTTCAAACACTCTTCAG
VastDB Features
Vast-tools module Information
Secondary ID
ENSGALG00000002451-29-37,29-36,29-35-1/3
Average complexity
Alt3
Mappability confidence:
NA
Protein Impact
ORF disruption when splice site is used (sequence exclusion)
No structure available
Features
Disorder rate (Iupred):
C1=0.000 A=NA C2=0.000
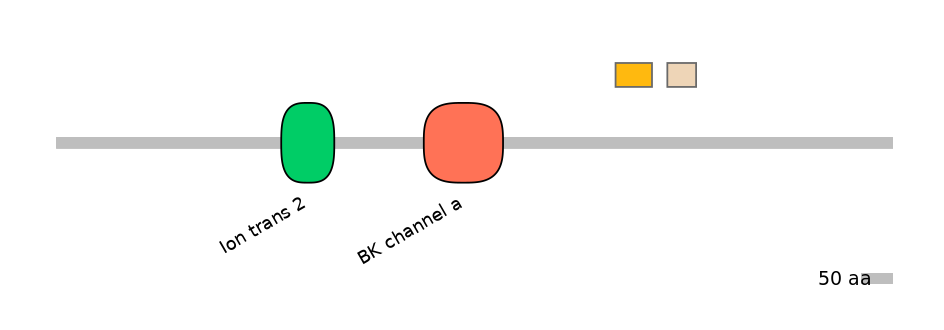
Domain overlap (PFAM):
C1:
NO
A:
NA
C2:
NO
Main Inclusion Isoform:
NA
Other Inclusion Isoforms:
NA
Other Skipping Isoforms:
NA
Associated events
Conservation
Mouse
(mm9)
No conservation detected
Rat
(rn6)
No conservation detected
Cow
(bosTau6)
No conservation detected
Zebrafish
(danRer10)
No conservation detected
Fruitfly
(dm6)
No conservation detected
Primers PCR
Suggestions for RT-PCR validation
F:
AGAAACTGCTGGAAATGGCTT
R:
TTCTTTGTCCACCACCACCAT
Band lengths:
128-161
Functional annotations
There are 0 annotated functions for this event
GENOMIC CONTEXT[edit]
INCLUSION PATTERN[edit]