MmuALTD0013315-1/2 @ mm9
Alternative 5'ss
Gene
ENSMUSG00000047793 | Sned1
Description
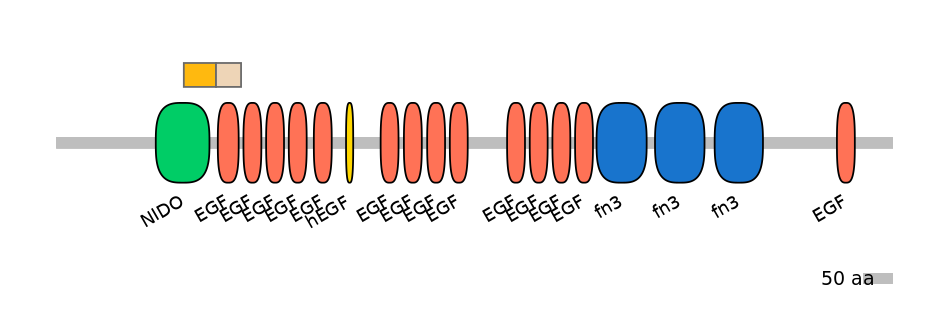
sushi, nidogen and EGF-like domains 1 [Source:MGI Symbol;Acc:MGI:3045960]
Coordinates
chr1:95156226-95158351:+
Coord C1 exon
chr1:95156226-95156356
Coord A exon
NA
Coord C2 exon
chr1:95158226-95158351
Length
0 bp
Sequences
Splice sites
5' ss Seq
CAGGTGCGC
5' ss Score
9.1
3' ss Seq
GCTCCTTCCTGTGTCCACAGCCT
3' ss Score
9.47
Exon sequences
Seq C1 exon
GTTTCAACGCAGGTGATGGGCACCGCTACTTCAACATCCCCGGGTCGCGCACAGCAGACATGGCTGAGGTGGAGACCACCACCAACGTGGGCGTGCCCGGCCGCTGGGCGTTTAGAATCGATGATGCCCAG
Seq A exon
NA
Seq C2 exon
CCTCTGTGTGCCTGGTCCTGCGTCCATGCCTCAATGGTGGCAAGTGCATTGATGACTGCGTCACGGGCAATCCCTCTTACACCTGTTCCTGTCTCGCTGGCTTCACAGGCCGGAGATGCCACCTGG
VastDB Features
Vast-tools module Information
Secondary ID
ENSMUSG00000047793-5-5,6-5-1/2
Average complexity
Alt5
Mappability confidence:
NA
Protein Impact
ORF disruption when splice site is used (sequence exclusion)
No structure available
Features
Disorder rate (Iupred):
C1=0.055 A=NA C2=0.000
Domain overlap (PFAM):
C1:
PF061199=NIDO=PD(47.3=78.2)
A:
NA
C2:
PF0000822=EGF=WD(100=83.7)
Main Inclusion Isoform:
NA
Other Inclusion Isoforms:
NA
Associated events
Other assemblies
Conservation
Human
(hg38)
No conservation detected
Rat
(rn6)
No conservation detected
Chicken
(galGal4)
No conservation detected
Zebrafish
(danRer10)
No conservation detected
Fruitfly
(dm6)
No conservation detected
Primers PCR
Suggestions for RT-PCR validation
F:
CAGACATGGCTGAGGTGGAGA
R:
ATGCACTTGCCACCATTGAGG
Band lengths:
126-154
Functional annotations
There are 0 annotated functions for this event
GENOMIC CONTEXT[edit]
INCLUSION PATTERN[edit]
SPECIAL DATASETS
- Pre-implantation embryo development
- Neural differentiation time course
- Muscular differentiation time course
- Spermatogenesis cell types
- Reprogramming of fibroblasts to iPSCs
- Hematopoietic precursors and cell types
Other AS DBs: