MmuEX0018452 @ mm9
Exon Skipping
Gene
ENSMUSG00000054942 | Fam73a
Description
family with sequence similarity 73, member A [Source:MGI Symbol;Acc:MGI:1924567]
Coordinates
chr3:151952020-151953873:-
Coord C1 exon
chr3:151953755-151953873
Coord A exon
chr3:151953424-151953496
Coord C2 exon
chr3:151952020-151952106
Length
73 bp
Sequences
Splice sites
3' ss Seq
CTCTGCTCTTCGTGATGCAGAAC
3' ss Score
7.56
5' ss Seq
CAGGTACTT
5' ss Score
8.17
Exon sequences
Seq C1 exon
CTCGCAGAGCAGAGAGAAGCGCAGCAGACCTTCAGCCTGGAGTCTTTTTGTCCCTGCCCTTTCTACGAAGAAGCTATGCATTTAGTTGAAGAAGGAAAAATTTACTCCAGAGTGCTGAG
Seq A exon
AACGGAGATGCTGGAGTGCCTGGGAGACAGTGACTTCCTCGCCAAACTTCACTGTATCCGGCAGGCTTTTCAG
Seq C2 exon
CTAATCCTTGCAGAAGCAGACAACAGGTCATTCCTTGCTGAAAGTGGAAGGAAAATTTTGTCAGCTTTAATTGTGAAAGCACGGAAG
VastDB Features
Vast-tools module Information
Secondary ID
ENSMUSG00000054942-'9-11,'9-10,10-11
Average complexity
C1
Mappability confidence:
100%=100=100%
Protein Impact
ORF disruption upon sequence exclusion
No structure available
Features
Disorder rate (Iupred):
C1=0.000 A=0.000 C2=0.000
Domain overlap (PFAM):
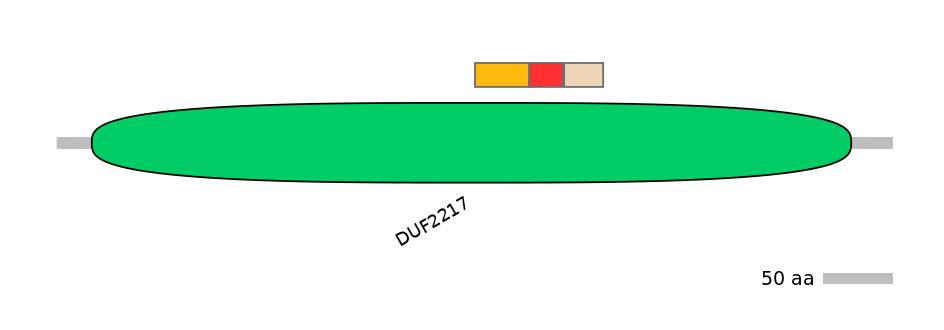
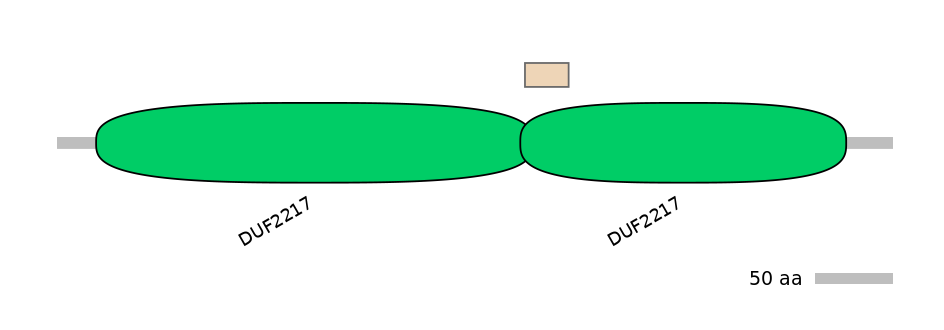
C1:
PF102654=DUF2217=FE(7.1=100)
A:
PF102654=DUF2217=FE(4.4=100)
C2:
PF102654=DUF2217=PD(1.8=17.2),PF048887=SseC=FE(28.6=100),PF102654=DUF2217=FE(13.3=100)
Other Inclusion Isoforms:
NA
Other Skipping Isoforms:
NA
Associated events
Other assemblies
Conservation
Rat
(rn6)
No conservation detected
Fruitfly
(dm6)
No conservation detected
Primers PCR
Suggestions for RT-PCR validation
F:
CAGAGCAGAGAGAAGCGCAG
R:
TCCTTCCACTTTCAGCAAGGA
Band lengths:
167-240
Functional annotations
There are 0 annotated functions for this event
GENOMIC CONTEXT[edit]
INCLUSION PATTERN[edit]
SPECIAL DATASETS
- Pre-implantation embryo development
- Neural differentiation time course
- Reprogramming of fibroblasts to iPSCs
- Hematopoietic precursors and cell types
Other AS DBs: