MmuEX0024743 @ mm9
Exon Skipping
Gene
ENSMUSG00000030287 | Itpr2
Description
inositol 1,4,5-triphosphate receptor 2 [Source:MGI Symbol;Acc:MGI:99418]
Coordinates
chr6:146366940-146371386:-
Coord C1 exon
chr6:146371228-146371386
Coord A exon
chr6:146369940-146369996
Coord C2 exon
chr6:146366940-146367038
Length
57 bp
Sequences
Splice sites
3' ss Seq
GTGTGAATTCAATTCCACAGGAC
3' ss Score
7.72
5' ss Seq
TAGGTAAGG
5' ss Score
9.31
Exon sequences
Seq C1 exon
CTGCTGCACATAAAAAGCAACAAGTACCTGACGGTGAACAAGAGGTTACCTGCCTTGCTGGAGAAGAATGCCATGCGAGTGTCACTGGATGCTGCAGGGAACGAAGGCTCCTGGTTCTACATCCATCCGTTCTGGAAGCTGAGAAGCGAGGGTGATAAT
Seq A exon
GACATGGGCGCCGTGATTTAATTGGGTTAATAAGTTTTCTGGTGATGTCTTCCTTAG
Seq C2 exon
ATCGTCGTAGGAGATAAAGTCGTTCTGATGCCTGTAAATGCTGGGCAGCCCCTGCATGCCAGCAACGTGGAGCTCTTGGACAACCCAGGCTGCAAAGAG
VastDB Features
Vast-tools module Information
Secondary ID
ENSMUSG00000030287_MULTIEX2-1/2=C1-2
Average complexity
S
Mappability confidence:
100%=100=100%
Protein Impact
ORF disruption upon sequence inclusion
No structure available
Features
Disorder rate (Iupred):
C1=0.000 A=0.000 C2=0.000
Domain overlap (PFAM):
C1:
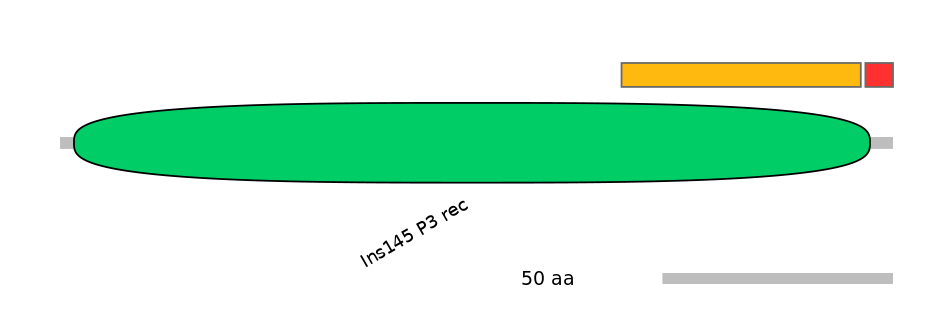
PF087096=Ins145_P3_rec=FE(23.0=100)
A:
PF087096=Ins145_P3_rec=PD(0.6=14.3)
C2:
PF087096=Ins145_P3_rec=FE(14.2=100)

Other Inclusion Isoforms:
NA
Associated events
Other assemblies
Conservation
Chicken
(galGal3)
No conservation detected
Zebrafish
(danRer10)
No conservation detected
Fruitfly
(dm6)
No conservation detected
Primers PCR
Suggestions for RT-PCR validation
F:
ACGAAGGCTCCTGGTTCTACA
R:
GTTGTCCAAGAGCTCCACGTT
Band lengths:
143-200
Functional annotations
There are 0 annotated functions for this event
GENOMIC CONTEXT[edit]
INCLUSION PATTERN[edit]
SPECIAL DATASETS
- Pre-implantation embryo development
- Neural differentiation time course
- Muscular differentiation time course
- Spermatogenesis cell types
- Reprogramming of fibroblasts to iPSCs
- Hematopoietic precursors and cell types
Other AS DBs: