RnoEX0067859 @ rn6
Exon Skipping
Gene
ENSRNOG00000053599 | Pogz
Description
pogo transposable element derived with ZNF domain [Source:RGD Symbol;Acc:1309980]
Coordinates
chr2:196021414-196023795:+
Coord C1 exon
chr2:196021414-196021522
Coord A exon
chr2:196023216-196023506
Coord C2 exon
chr2:196023586-196023795
Length
291 bp
Sequences
Splice sites
3' ss Seq
TTTGTCTCTTTTTTTAAAAGCCC
3' ss Score
7.65
5' ss Seq
TAGGTGAGA
5' ss Score
6.95
Exon sequences
Seq C1 exon
GGATTCCCTGTAAGGAATGTTCGCCCTGTACAGAATGCCATGAACCAGGTTGGGATCGTGCTGAATGTACAGCAAGGCCAAACAGTCAGACCAATTACACTGGTCCCAG
Seq A exon
CCCCAGGTACACAGTTTGTTAAGCCGACAGTTGGCGTCCCACAAGTGTTCTCTCAGATGACCCCCGTGAGGCCAGGCTCCACAATGCCTGTGAGGCCCACCACCAACACCTTTACCACCGTCATCCCAGCTACTCTCACCATCCGAAGCACTGTCCCACAGTCCCAGTCCCAGCAGACCAAGTCCACCCCCAGTACCTCCACTACTCCCATTGCCACACAGCCAACCGCATTGGGGCAGCTAGCTGGTCAGCCTCCAGGCCAGCCCAACCAGACCTCGAATCCCAAACTAG
Seq C2 exon
CTCCCTCCTTCCCCTCTCCACCTGCAGTGAGCATTGCCAGCTTTGTCACTGTGAAGCGACCTGGGGTTACAGGTGAAAATAGCAACGAAGTGGCCAAGTTGGTGAATACTCTTAACACTATCCCTTCCCTGGGCCAGAGTCCTGGGCCAGTGGTGGTGTCCAACAACAGTTCTGCTCAAAGAACCAGTGGACCCGAATCTTCAATGAAAG
VastDB Features
Vast-tools module Information
Secondary ID
ENSRNOG00000053599_MULTIEX1-1/2=C1-2
Average complexity
S
Mappability confidence:
100%=100=100%
Protein Impact
Alternative protein isoforms (Ref)
No structure available
Features
Disorder rate (Iupred):
C1=0.351 A=0.939 C2=0.944
Domain overlap (PFAM):
C1:
NO
A:
NO
C2:
NO
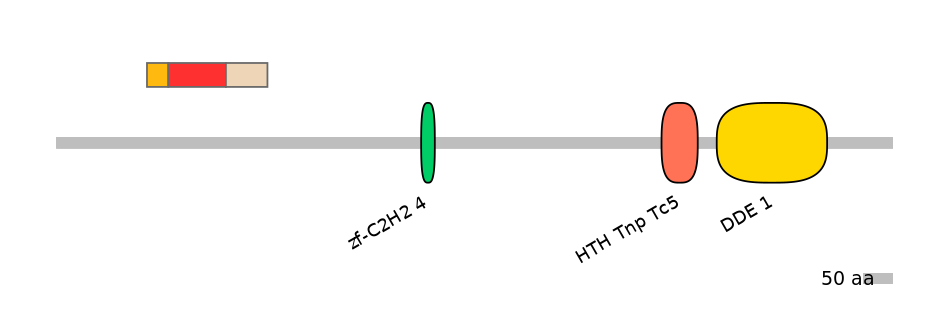
Main Skipping Isoform:
NA
Other Skipping Isoforms:
NA
Associated events
Conservation
Fruitfly
(dm6)
No conservation detected
Primers PCR
Suggestions for RT-PCR validation
F:
CCCTGTAAGGAATGTTCGCCC
R:
TGAAGATTCGGGTCCACTGGT
Band lengths:
307-598
Functional annotations
There are 0 annotated functions for this event
GENOMIC CONTEXT[edit]
INCLUSION PATTERN[edit]