GgaEX6042254 @ galGal4
Exon Skipping
Gene
ENSGALG00000009287 | ATF2
Description
Cyclic AMP-dependent transcription factor ATF-2 [Source:UniProtKB/Swiss-Prot;Acc:O93602]
Coordinates
chr7:16173365-16179779:+
Coord C1 exon
chr7:16173365-16173493
Coord A exon
chr7:16179194-16179372
Coord C2 exon
chr7:16179665-16179779
Length
179 bp
Sequences
Splice sites
3' ss Seq
GATCTTTGGTTTCTCCTCAGGAA
3' ss Score
9.34
5' ss Seq
AGTGTAAGT
5' ss Score
8.46
Exon sequences
Seq C1 exon
ATGCCTCTAGATCTGTCTCCTCTTGCAACACCAATTATAAGAAACAAAATTGAAGAACCTTCTGTTGTAGAGACGACTCATCAAGATAGTCCTTTACCTCATCCAGAGTCTACTACCAATGATGAAAAG
Seq A exon
GAAGTGTCCCTGCAGCAGACTGCACAGCCCACATCAACAATAGTTCGTCCAGCATCGCTACAGGTTCCTAACGTACTACTTACAAGTTCTGACTCAAGTGTTATTATTCAGCAGGCAATACCATCACCAACGTCTAGTACTGTTATTACCCAGGCTCCATCCTCTAACAGACCAATAGT
Seq C2 exon
TCCAGTTCCAGGCCCGTTTCCTCTGCTGTTACATCTTCCTAATGGACAAACCATGCCTGTTGCTATTCCTGCATCTATTACAAATTCCAACGTGCATGTCCCTGCTGCAGTTCCA
VastDB Features
Vast-tools module Information
Secondary ID
ENSGALG00000009287-'10-19,'10-15,13-19=AN
Average complexity
A_S
Mappability confidence:
100%=100=100%
Protein Impact
ORF disruption upon sequence exclusion
No structure available
Features
Disorder rate (Iupred):
C1=0.890 A=0.583 C2=0.923
Domain overlap (PFAM):
C1:
NO
A:
NO
C2:
NO
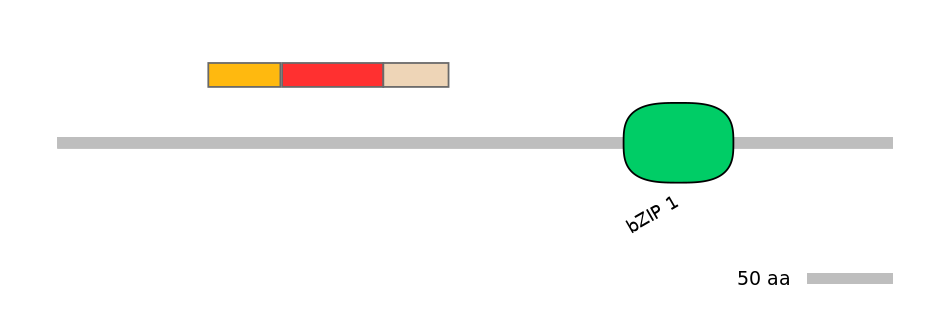
Main Skipping Isoform:
ENSGALT00000033263fB1563
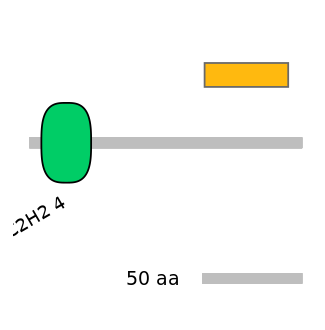
Other Skipping Isoforms:
NA
Associated events
Other assemblies
Conservation
Fruitfly
(dm6)
No conservation detected
Primers PCR
Suggestions for RT-PCR validation
F:
TGCCTCTAGATCTGTCTCCTCT
R:
GGAACTGCAGCAGGGACATG
Band lengths:
242-421
Functional annotations
There are 1 annotated functions for this event
PMID: 26538579
Encodes an experimentally validated Eukaryotic Linear Motif (ELM). Method: protein array. ELM ID: ELMI003089; ELM sequence: RPASLQV; Overlap: FULL
GENOMIC CONTEXT[edit]
INCLUSION PATTERN[edit]