HsaEX6101504 @ hg19
Exon Skipping
Gene
ENSG00000113240 | CLK4
Description
CDC-like kinase 4 [Source:HGNC Symbol;Acc:13659]
Coordinates
chr5:178044345-178050417:-
Coord C1 exon
chr5:178050257-178050417
Coord A exon
chr5:178045557-178045779
Coord C2 exon
chr5:178044345-178044435
Length
223 bp
Sequences
Splice sites
3' ss Seq
ATCTTTTAATGTGTTTGCAGTCA
3' ss Score
6.59
5' ss Seq
TCGGTATGA
5' ss Score
7.72
Exon sequences
Seq C1 exon
ATGCGGCATTCCAAAAGAACTCACTGTCCTGATTGGGATAGCAGAGAAAGCTGGGGACATGAAAGCTATCGTGGAAGTCACAAGCGGAAGAGGAGATCTCATAGTAGCACACAAGAGAACAGGCATTGTAAACCACATCACCAGTTTAAAGAATCTGATTG
Seq A exon
TCATTATTTAGAAGCAAGGTCCTTGAATGAGCGAGATTATCGGGACCGGAGATACGTTGACGAATACAGGAATGACTACTGTGAAGGATATGTTCCTAGACATTATCACAGAGACATTGAAAGCGGGTATCGAATCCACTGCAGTAAATCTTCAGTCCGCAGCAGGAGAAGCAGTCCTAAAAGGAAGCGCAATAGACACTGTTCAAGTCATCAGTCACGTTCG
Seq C2 exon
AAGAGCCACCGAAGGAAAAGATCCAGGAGTATAGAGGATGATGAGGAGGGTCACCTGATCTGTCAAAGTGGAGACGTTCTAAGAGCAAGAT
VastDB Features
Vast-tools module Information
Secondary ID
ENSG00000113240-'2-17,'2-16,9-17=AN
Average complexity
A_C2
Mappability confidence:
100%=100=100%
Protein Impact
ORF disruption upon sequence exclusion
No structure available
Features
Disorder rate (Iupred):
C1=0.459 A=0.329 C2=0.323
Domain overlap (PFAM):
C1:
NO
A:
NO
C2:
PF0006920=Pkinase=PU(0.1=0.0)
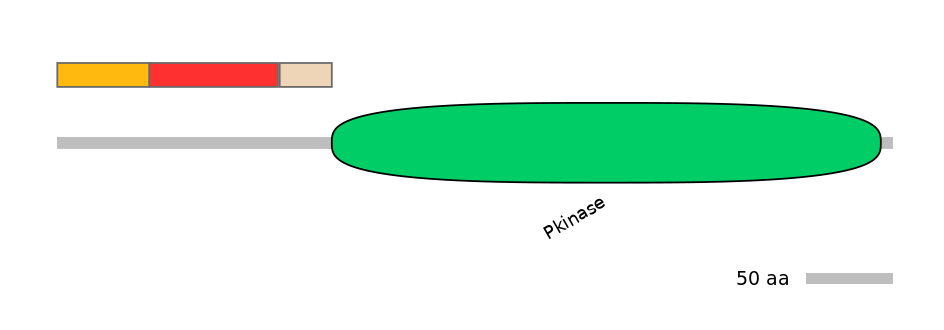
Other Skipping Isoforms:
NA
Associated events
Other assemblies
Conservation
Fruitfly
(dm6)
No conservation detected
Primers PCR
Suggestions for RT-PCR validation
F:
ATGCGGCATTCCAAAAGAACT
R:
ATCTTGCTCTTAGAACGTCTCCAC
Band lengths:
252-475
Functional annotations
There are 0 annotated functions for this event
GENOMIC CONTEXT[edit]
INCLUSION PATTERN[edit]
SPECIAL DATASETS
- Autistic and control brains
Other AS DBs:
FasterDB (Includes CLIP-seq data)
AS-ALPS (AS-induced ALteration of Protein Structure, links to PINs)
APPRIS (Selection of principal isoform)
DEU primates (Only for human)