MmuALTA0002351-3/3 @ mm9
Alternative 3'ss
Gene
ENSMUSG00000019302 | Atp6v0a1
Description
ATPase, H+ transporting, lysosomal V0 subunit A1 [Source:MGI Symbol;Acc:MGI:103286]
Coordinates
chr11:100882368-100888077:+
Coord C1 exon
chr11:100882368-100882496
Coord A exon
chr11:100887974-100887994
Coord C2 exon
chr11:100887995-100888077
Length
21 bp
Sequences
Splice sites
5' ss Seq
GAGGTCAGG
5' ss Score
4.94
3' ss Seq
TAATTGATTTGTGAATTCAGGCT
3' ss Score
2.8
Exon sequences
Seq C1 exon
GCCAATTTCGAGAAGATTGAAAATGAACTGAAGGAAATCAACACTAACCAGGAAGCTCTGAAGAGAAACTTCCTGGAACTGACTGAATTAAAATTTATCCTGCGAAAAACCCAGCAGTTTTTCGATGAG
Seq A exon
GCTGAATTGCATCATCAGCAG
Seq C2 exon
ATGGCGGATCCAGACCTGTTGGAAGAGTCCTCATCACTCTTGGAGCCAAACGAGATGGGAAGAGGCGCACCCTTACGACTTGG
VastDB Features
Vast-tools module Information
Secondary ID
ENSMUSG00000019302-6-10,6-9,6-8-3/3
Average complexity
Alt3
Mappability confidence:
NA
Protein Impact
Protein isoform when splice site is used (No Ref)
No structure available
Features
Disorder rate (Iupred):
C1=0.000 A=NA C2=0.239
Domain overlap (PFAM):
C1:
PF0149614=V_ATPase_I=FE(5.2=100)
A:
NA
C2:
PF0149614=V_ATPase_I=FE(3.4=100)
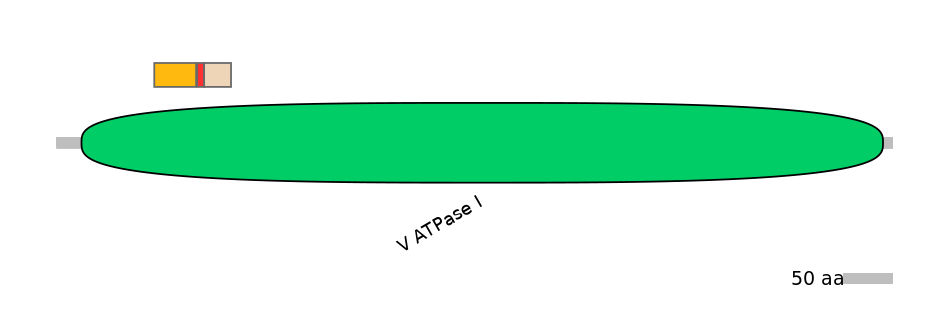
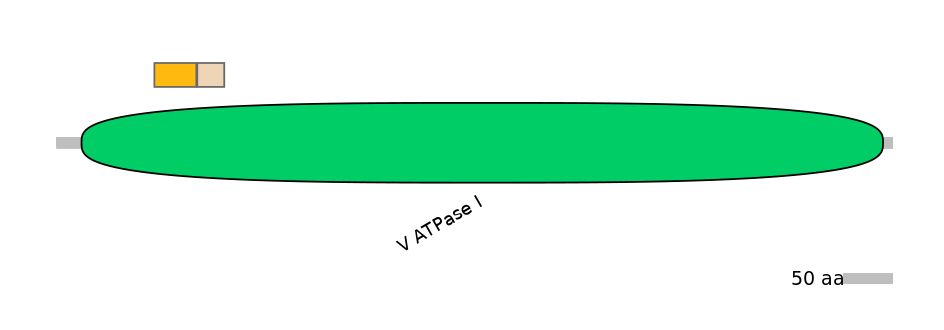
Other Inclusion Isoforms:
NA
Associated events
Other assemblies
Conservation
Zebrafish
(danRer10)
No conservation detected
Fruitfly
(dm6)
No conservation detected
Primers PCR
Suggestions for RT-PCR validation
F:
TTATCCTGCGAAAAACCCAGCA
R:
CGCCTCTTCCCATCTCGTTTG
Band lengths:
102-123
Functional annotations
There are 0 annotated functions for this event
GENOMIC CONTEXT[edit]
INCLUSION PATTERN[edit]
SPECIAL DATASETS
- Pre-implantation embryo development
- Neural differentiation time course
- Muscular differentiation time course
- Spermatogenesis cell types
- Reprogramming of fibroblasts to iPSCs
- Hematopoietic precursors and cell types
Other AS DBs: