MmuEX0041329 @ mm9
Exon Skipping
Gene
ENSMUSG00000016763 | Scube1
Description
signal peptide, CUB domain, EGF-like 1 [Source:MGI Symbol;Acc:MGI:1890616]
Coordinates
chr15:83469134-83489581:-
Coord C1 exon
chr15:83489456-83489581
Coord A exon
chr15:83484599-83484715
Coord C2 exon
chr15:83469134-83469250
Length
117 bp
Sequences
Splice sites
3' ss Seq
TACACTTTGGGACTAACTAGTAA
3' ss Score
0.3
5' ss Seq
TCGGTAAGT
5' ss Score
11.78
Exon sequences
Seq C1 exon
AGGGTATGAACTGCATGAACAAAGACCATGGCTGTGCCCACATCTGCCGGGAGACGCCCAAAGGTGGGGTAGCCTGTGACTGCAGGCCCGGCTTTGACCTTGCCCAAAACCAGAAGGACTGCACAC
Seq A exon
TAACCTGTAACTATGGAAACGGAGGCTGTCAGCACAGCTGCGAGGACACGGACACGGGCCCCATGTGTGGCTGCCACCAGAAGTACGCCTTACATGCAGATGGTCGGACGTGCATCG
Seq C2 exon
AGACATGTGCAGTGAACAACGGAGGCTGTGACCGGACGTGTAAGGACACTGCCACTGGCGTGCGGTGCAGCTGCCCGGTTGGATTCACGCTGCAGCCAGATGGAAAGACATGCAAAG
VastDB Features
Vast-tools module Information
Secondary ID
ENSMUSG00000016763_MULTIEX1-4/9=3-6
Average complexity
C1
Mappability confidence:
100%=100=100%
Protein Impact
Alternative protein isoforms (Ref)
No structure available
Features
Disorder rate (Iupred):
C1=0.000 A=0.000 C2=0.000
Domain overlap (PFAM):
C1:
PF146701=FXa_inhibition=WD(100=86.0)
A:
PF146701=FXa_inhibition=WD(100=90.0)
C2:
PF146701=FXa_inhibition=WD(100=90.0)
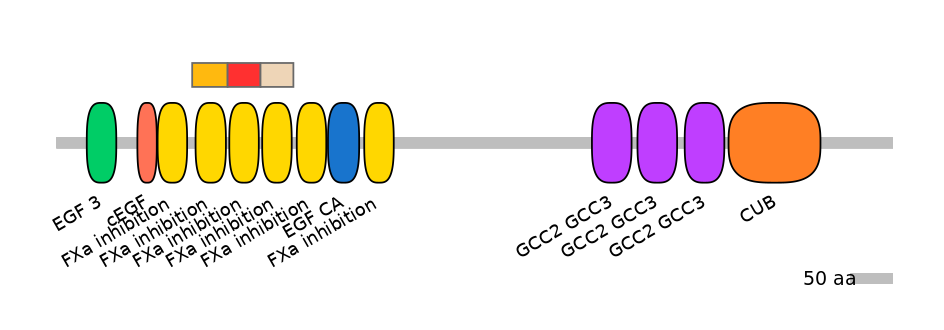
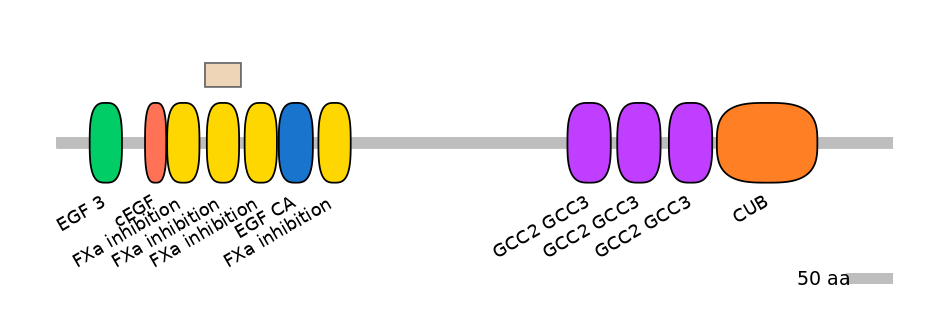
Other Skipping Isoforms:
NA
Associated events
Other assemblies
Conservation
Fruitfly
(dm6)
No conservation detected
Primers PCR
Suggestions for RT-PCR validation
F:
AGGGTATGAACTGCATGAACA
R:
TTTGCATGTCTTTCCATCTGGCT
Band lengths:
242-359
Functional annotations
There are 0 annotated functions for this event
GENOMIC CONTEXT[edit]
INCLUSION PATTERN[edit]
SPECIAL DATASETS
- Pre-implantation embryo development
- Neural differentiation time course
- Muscular differentiation time course
- Spermatogenesis cell types
- Reprogramming of fibroblasts to iPSCs
- Hematopoietic precursors and cell types
Other AS DBs: