RnoEX0102023 @ rn6
Exon Skipping
Gene
ENSRNOG00000003707 | Zmym3
Description
zinc finger MYM-type containing 3 [Source:RGD Symbol;Acc:1564967]
Coordinates
chrX:71308798-71309518:-
Coord C1 exon
chrX:71309368-71309518
Coord A exon
chrX:71309043-71309158
Coord C2 exon
chrX:71308798-71308884
Length
116 bp
Sequences
Splice sites
3' ss Seq
TCTTTCTTTCTATGCCCCAGGCC
3' ss Score
9.58
5' ss Seq
CAGGTACTC
5' ss Score
7.04
Exon sequences
Seq C1 exon
AAAAACACACGTGTGTACCCATGTGTCTGGTGCAAGACCCTGTGTAAGAACTTTGAGATGCTATCACATGTGGATCGTAATGGCAAGACCAGCTTGTTCTGTTCCCTGTGCTGTACCACTTCTTACAAAGTGAAGCAGGCAGGGCTCACTG
Seq A exon
GCCCTCCCCGACCCTGCAGCTTCTGCCGCCGCAGCCTCTCTGACCCTTGTTACTACAACAAAGTTGATCGCACAGTCTACCAGTTCTGCAGCCCCAGCTGCTGGACCAAGTTCCAG
Seq C2 exon
CATACTAGCCCTGAGGGGGGCATTCACCTGAGCTGTCACTACTGCCATAGCCTCTTCAGTGGCAAGCCTGAGGTCTTGGAGTGGCAG
VastDB Features
Vast-tools module Information
Secondary ID
ENSRNOG00000003707-'15-16,'15-15,16-16
Average complexity
S
Mappability confidence:
100%=100=100%
Protein Impact
ORF disruption upon sequence exclusion
No structure available
Features
Disorder rate (Iupred):
C1=0.000 A=0.000 C2=0.000
Domain overlap (PFAM):
C1:
PF064679=zf-FCS=PD(95.3=80.4),PF064679=zf-FCS=PU(7.3=5.9)
A:
PF064679=zf-FCS=PD(90.2=94.9)
C2:
PF064679=zf-FCS=PU(61.0=86.2)
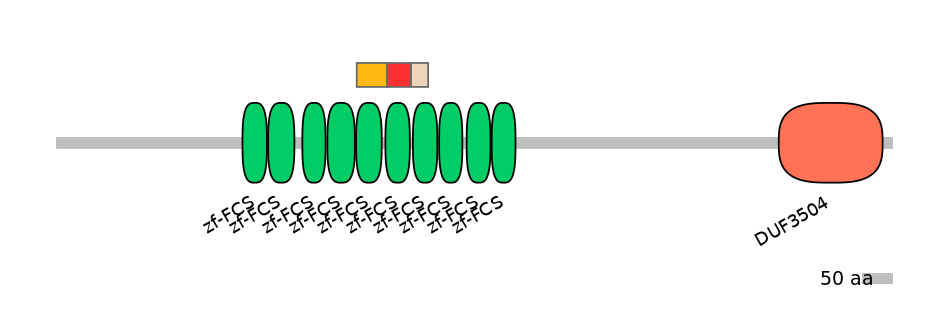
Main Skipping Isoform:
NA
Other Inclusion Isoforms:
NA
Other Skipping Isoforms:
NA
Associated events
Conservation
Fruitfly
(dm6)
No conservation detected
Primers PCR
Suggestions for RT-PCR validation
F:
CACACGTGTGTACCCATGTGT
R:
CACTCCAAGACCTCAGGCTTG
Band lengths:
229-345
Functional annotations
There are 0 annotated functions for this event
GENOMIC CONTEXT[edit]
INCLUSION PATTERN[edit]