DreEX0055532 @ danRer10
Exon Skipping
Gene
ENSDARG00000086927 | pik3c2b
Description
phosphatidylinositol-4-phosphate 3-kinase, catalytic subunit type 2 beta [Source:ZFIN;Acc:ZDB-GENE-050309-145]
Coordinates
chr11:23362060-23363853:-
Coord C1 exon
chr11:23363684-23363853
Coord A exon
chr11:23362338-23362494
Coord C2 exon
chr11:23362060-23362169
Length
157 bp
Sequences
Splice sites
3' ss Seq
CTTTTTTTTTTTTTTTTCAGCGA
3' ss Score
11.53
5' ss Seq
CTGGTGAGT
5' ss Score
10.1
Exon sequences
Seq C1 exon
GCGGTGGAGAACTTCATCTATTCGTGCGCTGGTTGTTGTGTGGCCACATACATTCTGGGCATCTGTGACCGGCACAATGACAACATCATGCTGAAGACCAGCGGACACATGTTCCACATTGACTTTGGCAAGTTCCTGGGGCACGCACAGATGTTTGGCAACATTAAACG
Seq A exon
CGACCGGGCTCCTTTTGTGTTTACATCTGACATGGCGTATGTCATCAATGGAGGTGACAAGCCATCCAGTCGTTTTCATGATTTTGTAGATTTGTGTTGTGAGGCCTACAATCGCATCAGGAAACACGCACATCTGTTTCTCAACCTGCTCGGTCTG
Seq C2 exon
ATGCTGTCCAGTGGAATACCAGAGCTGTCTGATGTGGATGATCTGAAGTATGTCTATGATGCACTGAGGCCACATGAATCAGAAGCTGATGCCACTATGCACTTCACCAG
VastDB Features
Vast-tools module Information
Secondary ID
ENSDARG00000086927-'33-31,'33-30,34-31
Average complexity
S
Mappability confidence:
100%=100=100%
Protein Impact
ORF disruption upon sequence exclusion
No structure available
Features
Disorder rate (Iupred):
C1=0.000 A=0.000 C2=0.000
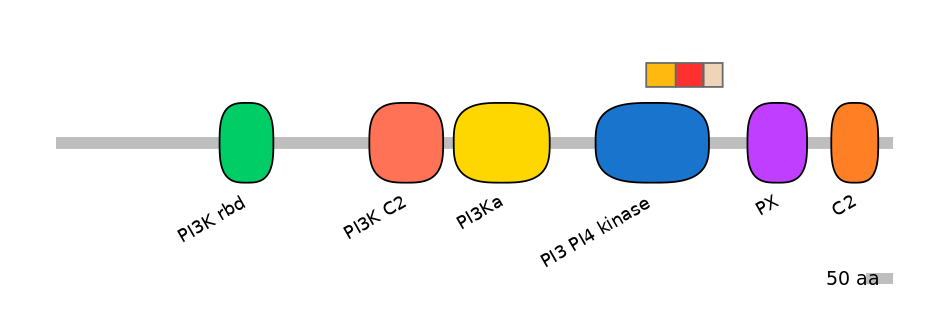
Domain overlap (PFAM):
C1:
PF0045422=PI3_PI4_kinase=FE(25.9=100)
A:
PF0045422=PI3_PI4_kinase=FE(24.1=100)
C2:
PF0045422=PI3_PI4_kinase=PD(4.6=27.0)
Main Skipping Isoform:
NA
Other Inclusion Isoforms:
NA
Other Skipping Isoforms:
NA
Associated events
Conservation
Fruitfly
(dm6)
No conservation detected
Primers PCR
Suggestions for RT-PCR validation
F:
TGTGGCCACATACATTCTGGG
R:
CTGGTGAAGTGCATAGTGGCA
Band lengths:
242-399
Functional annotations
There are 0 annotated functions for this event
GENOMIC CONTEXT[edit]
INCLUSION PATTERN[edit]