GgaEX1000566 @ galGal3
Exon Skipping
Description
NA
Coordinates
chr23:4685327-4687885:-
Coord C1 exon
chr23:4687633-4687885
Coord A exon
chr23:4685654-4685828
Coord C2 exon
chr23:4685327-4685539
Length
175 bp
Sequences
Splice sites
3' ss Seq
TTGTTTTTCTTTTCCTGTAGAGA
3' ss Score
10.47
5' ss Seq
GTGGTAATG
5' ss Score
4.46
Exon sequences
Seq C1 exon
CTCCACAGCTGTCATCAGGTTTCCAGCCTGCTGTGGCATCCTCTGGCATGAGTAAAATGCTTCCTTCAGTTCCAGGCACAGCTGTTCGGGTTTCCTGTTCTGGCTGTAAAAAAATCCTCCAGAAGGGGCAGACCGCATACCAGAGGAAAGGCTCTACCCAGCTCTTCTGCTCCACACTGTGCCTCACTGGATACACCGTTCCAGCTTGTCGCCCACCAGCTTCCACCAAGAAAACCTGCTCAAGCTGCTCGAA
Seq A exon
AGAAATTCTAAATCCAAAGGATGTAATCACTGCCCAGTTTGACAATACAAATTCCAGTAAGGATTTCTGCAGCCAGTCATGTCTTTCTACGTATGAAATGAAAAGGAAACCTATTATTACTATTCACACTAACAGCATTTCAACCAAGTGCAGCATGTGCCAGAAGAATGCAGTG
Seq C2 exon
ATCAGACATGAAGTGAATTACCAGAACGTCGTACACAAGCTCTGCAGTGATGCCTGCTTCTCCAAGTTCCGCTCTGCTAATAACTTAACCATGAACTGCTGTGAGAATTGTGGAGGTTACTGCTACAGTGGGTCTGGGCAGTGTCGCATGCTGCAGATAGAGGGACAGTCAAAAAAGTTCTGCAGTTCGACATGTGTGACACTGTACAAGCAG
VastDB Features
Vast-tools module Information
Secondary ID
ENSGALG00000002499-'10-10,'10-9,11-10=AN
Average complexity
A_S
Mappability confidence:
100%=100=100%
Protein Impact
ORF disruption upon sequence exclusion
No structure available
Features
Disorder rate (Iupred):
C1=0.106 A=0.000 C2=0.000
Domain overlap (PFAM):
C1:
PF064679=zf-FCS=WD(100=48.2),PF064679=zf-FCS=PU(28.9=15.3)
A:
PF064679=zf-FCS=PD(68.9=52.5),PF064679=zf-FCS=PU(40.0=27.1)
C2:
PF064679=zf-FCS=PD(55.0=31.0),PF064679=zf-FCS=WD(100=62.0),PF064679=zf-FCS=PU(0.1=0.0)
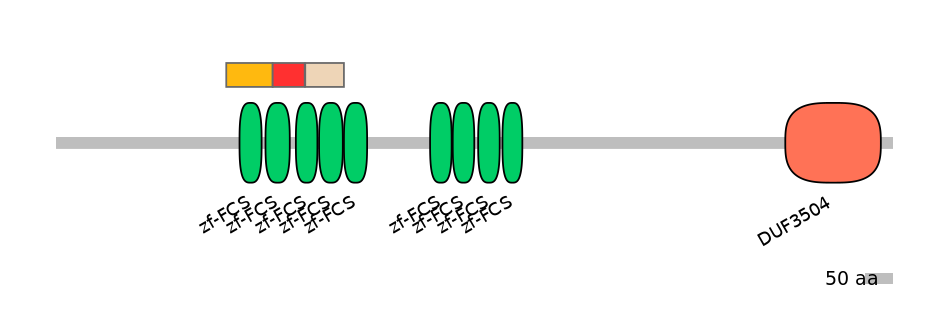
Main Skipping Isoform:
NA
Other Skipping Isoforms:
NA
Associated events
Other assemblies
Conservation
Fruitfly
(dm6)
No conservation detected
Primers PCR
Suggestions for RT-PCR validation
F:
AGGCTCTACCCAGCTCTTCTG
R:
CCCAGACCCACTGTAGCAGTA
Band lengths:
243-418
Functional annotations
There are 0 annotated functions for this event
GENOMIC CONTEXT[edit]
INCLUSION PATTERN[edit]