GgaEX1043338 @ galGal3
Exon Skipping
Gene
ENSGALG00000004029 | PHF12
Description
NA
Coordinates
chr19:5852197-5855866:-
Coord C1 exon
chr19:5855705-5855866
Coord A exon
chr19:5853867-5854683
Coord C2 exon
chr19:5852197-5852319
Length
817 bp
Sequences
Splice sites
3' ss Seq
TTTTTCTCCCTTCCTCCCAGTGG
3' ss Score
10.06
5' ss Seq
CAGGTAAGT
5' ss Score
10.86
Exon sequences
Seq C1 exon
GTTCCTGATGCTATAAAGTCCCAATACCGGCTTCCACCCCCGTTACTTGCGCCTGCAGCAATTAGAGATGGCGAGCTGATCTGTAACGGGATCCCTGAGGAACCCTTGCAGAAGCACCTTTTGAACACTGAGCACTTAGCCAGCCAGACAGAACAACAGGAG
Seq A exon
TGGCTCTGTAGTGTTGTTGCGCTCCAGTGCAGCATATTGAAACATTTATCTGCTAAGCAGATGACTTCGCTTTGGGACTCTGAACAAACAGAGAAGGCTGATATTAAGCCTGTTATTGTGGCAGACGGCACGCTCGCCAACCCTTACCAGGCAGCTGACAAGGCACCCACACCTTCCCTCTATTCCATGTCATCCTGCACCTCAGGGATTACCACCCAGAATTCCTTGAGTTCCTCACAATCCCAGCAGACTTTGTCACAGGAAGATGTCAACTGCAGCTCCTGTCTGGATAAATCCAAGAAGGTGTCTTCTTGCGGGACTTCAAATGGGCCGACGGCCTCTGACGTTAAAGTAAATGGTCCTCATTTCTATGGCGTTTCATCCGAACCTTCAGCACAGTTGACTGACTCCCAGAGGCTGATTGGAGCTGGTAACACAATACCTGTTTTGTCTCATCGGCAAACGTGGCCCAGGCCCCTCACGCCCCCAGCGACTTGTGGACTTCTGAATCACACCATTGGAACCATTGTCAAAACGGAGAACGTAGGAGGACCCGTTTCTTGTGCACAAAGGGGTTCTGTGCCAGTCACAAGTATGCCTACTTCCATATCGAGCTCCTGTGCCAGCCTGGAGACCACCAGCACTTTGCAAAGGAAGAATGTGCAGACGCAGATAGGGCCGCCGCTGTCAGCAGACCTGCGATCCATGGGCTCTCCTCTGACTGCCACCAGGGCTCTCACCCCTCCCCAGGCTGCAGGAGATGGGATCTCGGCTGTTGGATCCGCAAACAGATTCTGTTCACCAGCACCACCTTCAG
Seq C2 exon
ATGGTAAGGTCAGCCCCAGCACCTTATCCATAGGAAGCGCTTTAACTCTACCCTCATCGCTCACTTCAAACTCTGCAGCTATGTTGGACCTCACCAGTTCCCTGAAAGCGTTTATGGATGGCG
VastDB Features
Vast-tools module Information
Secondary ID
ENSGALG00000004029-'10-18,'10-14,12-18=AN
Average complexity
A_S
Mappability confidence:
100%=100=100%
Protein Impact
ORF disruption upon sequence exclusion
No structure available
Features
Disorder rate (Iupred):
C1=0.111 A=0.511 C2=0.214
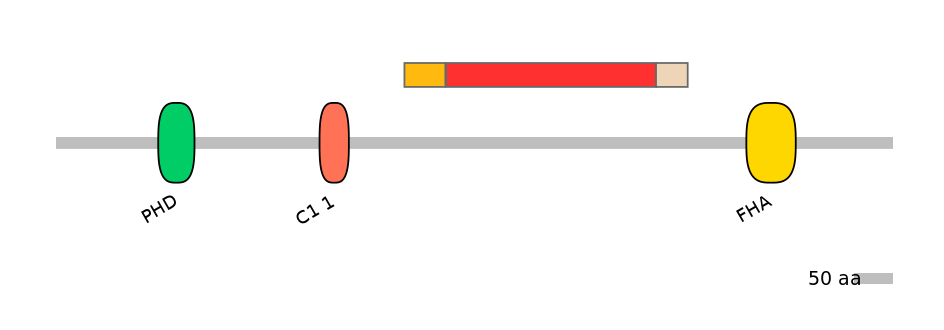
Domain overlap (PFAM):
C1:
NO
A:
NO
C2:
NO
Main Skipping Isoform:
NA
Other Skipping Isoforms:
NA
Associated events
Other assemblies
Conservation
Fruitfly
(dm6)
No conservation detected
Primers PCR
Suggestions for RT-PCR validation
F:
ATGCTATAAAGTCCCAATACCGGC
R:
CGCCATCCATAAACGCTTTCA
Band lengths:
278-1095
Functional annotations
There are 2 annotated functions for this event
PMID: 24782567
Encodes an experimentally validated Eukaryotic Linear Motif (ELM). Method: detection by mass spectrometry. ELM ID: ELMI002883; ELM sequence: EKADIKPV; Overlap: FULL
PMID: 25218447
Encodes an experimentally validated Eukaryotic Linear Motif (ELM). Method: Identification by mass spectrometry, detection by mass spectrometry, his tag, pull down. ELM ID: ELMI002883; ELM sequence: EKADIKPV; Overlap: FULL
GENOMIC CONTEXT[edit]
INCLUSION PATTERN[edit]