HsaEX6017546 @ hg38
Exon Skipping
Gene
ENSG00000118997 | DNAH7
Description
dynein axonemal heavy chain 7 [Source:HGNC Symbol;Acc:HGNC:18661]
Coordinates
chr2:196048296-196058116:-
Coord C1 exon
chr2:196058054-196058116
Coord A exon
chr2:196051187-196051249
Coord C2 exon
chr2:196048296-196048404
Length
63 bp
Sequences
Splice sites
3' ss Seq
CCTTCATATTTTTCACACAGGAG
3' ss Score
8.03
5' ss Seq
ATGGTAACT
5' ss Score
6.06
Exon sequences
Seq C1 exon
GATAAATCGGCCAGCAAAGAAAAATCCAAGAAACCAGTAAGATTTCTACCACAGCTGTCTATG
Seq A exon
GAGAAATTAGCCAGCAAAGAAAAGTTCAAGGCACCAGCAAGAGCTTTACCACAGCTGTCTATG
Seq C2 exon
GTGAGTACAAAGCCCCACTGGCAGCAGGCAGCTCCATCATTCCATTTGAGTGTAAAGCAGGATGATGAGAGTCCAGAACCATTTAGTGTTAAAAATGAACAGTCCCATG
VastDB Features
Vast-tools module Information
Secondary ID
ENSG00000118997-'4-5,'4-3,7-5
Average complexity
C1
Mappability confidence:
100%=100=100%
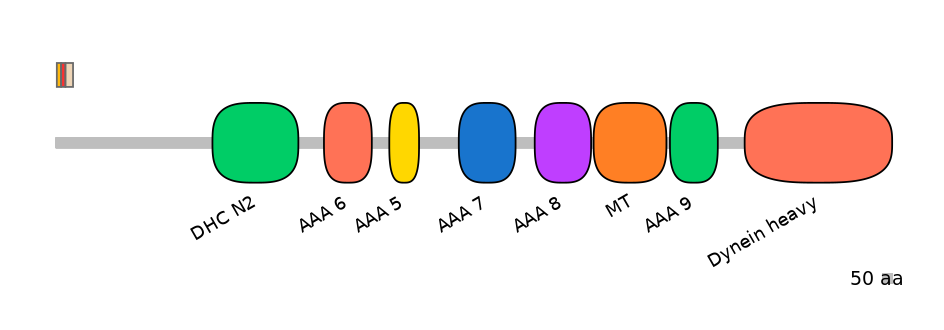
Protein Impact
Alternative protein isoforms (Ref)
No structure available
Features
Disorder rate (Iupred):
C1=1.000 A=0.915 C2=0.973
Domain overlap (PFAM):
C1:
NO
A:
NO
C2:
NO
Main Skipping Isoform:
NA
Other Skipping Isoforms:
NA
Associated events
Other assemblies
Conservation
Mouse
(mm9)
No conservation detected
Chicken
(galGal4)
No conservation detected
Chicken
(galGal3)
No conservation detected
Zebrafish
(danRer10)
No conservation detected
Fruitfly
(dm6)
No conservation detected
Primers PCR
Suggestions for RT-PCR validation
F:
CGGCCAGCAAAGAAAAATCCA
R:
TCTGGACTCTCATCATCCTGCT
Band lengths:
133-196
Functional annotations
There are 0 annotated functions for this event
GENOMIC CONTEXT[edit]
INCLUSION PATTERN[edit]
SPECIAL DATASETS
- Autistic and control brains