MmuEX6028596 @ mm10
Exon Skipping
Gene
ENSMUSG00000015829 | Tnr
Description
tenascin R [Source:MGI Symbol;Acc:MGI:99516]
Coordinates
chr1:159886057-159888376:+
Coord C1 exon
chr1:159886057-159886320
Coord A exon
chr1:159886870-159887139
Coord C2 exon
chr1:159888257-159888376
Length
270 bp
Sequences
Splice sites
3' ss Seq
CAATCTTTTCTACTTTGCAGGCT
3' ss Score
8.27
5' ss Seq
CAGGTGAGG
5' ss Score
10.07
Exon sequences
Seq C1 exon
AACTTGACAGTCCCCGAGACCTCATGGTAACAGCCTCCTCAGAGACCTCTATCTCTCTCATCTGGACCAAGGCCAGTGGTCCCATTGATCACTACAGAATTACTTTTACCCCATCTTCTGGGATCTCCTCCGAAGTCACTGTGCCCAGGGATAGGACTTCATACACACTAACAGATCTAGAGCCTGGAGCAGAATACATTATCTCCATCACTGCTGAGAGGGGTCGGCAGCAGAGCCTGGAGTCTACTGTGGATGCCTTCACAG
Seq A exon
GCTTCCGCCCTATCTCCCATTTGCACTTTTCTCATGTGACCTCCTCCAGTGTCAATATCACCTGGAGTGACCCATCTCCCCCAGCAGACAGACTCATTCTGAACTACAGCCCCAGGGACAAAGAGGAAGACATGTTGGAGGTCCTCTTGGATGCCACCAAGAGGCACGCTGTCTTGATGGGTCTACAGCCAGCCACTGAATATATAGTGAACCTTGTAGCTGTCCATGGGACGGTAACCTCTGAACCCATAGTGGGTTCTATCACTACAG
Seq C2 exon
GAATTGATCCTCCCAAAAACATCACAATTAGCAACGTGACTAAGGACTCCCTGACGGTGTCCTGGAGCTCTCCTGTTGCGCCTTTTGATTACTACCGAGTATCGTACCGACCCACCCAAG
VastDB Features
Vast-tools module Information
Secondary ID
ENSMUSG00000015829-'27-22,'27-21,28-22
Average complexity
S
Mappability confidence:
100%=100=100%
Protein Impact
Alternative protein isoforms (Ref)
No structure available
Features
Disorder rate (Iupred):
C1=0.247 A=0.088 C2=0.024
Domain overlap (PFAM):
C1:
PF0004116=fn3=WD(100=89.9)
A:
PF0004116=fn3=WD(100=89.0)
C2:
PF0004116=fn3=PU(47.5=92.7)
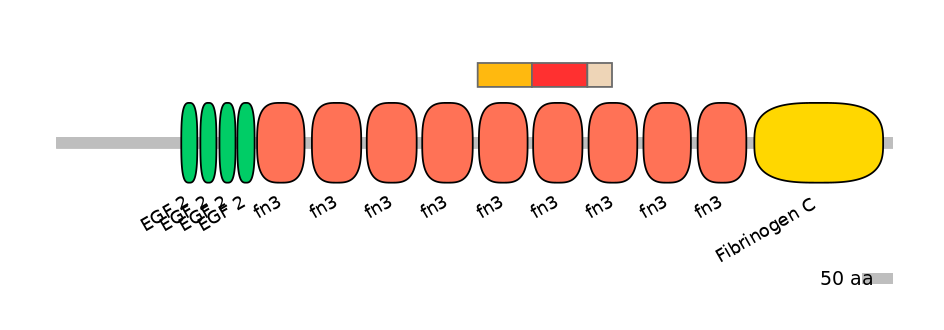
Main Skipping Isoform:
NA
Other Skipping Isoforms:
NA
Associated events
Other assemblies
Conservation
Fruitfly
(dm6)
No conservation detected
Primers PCR
Suggestions for RT-PCR validation
F:
AACAGCCTCCTCAGAGACCTC
R:
CACCGTCAGGGAGTCCTTAGT
Band lengths:
295-565
Functional annotations
There are 0 annotated functions for this event
GENOMIC CONTEXT[edit]
INCLUSION PATTERN[edit]
SPECIAL DATASETS
- Ribosome-engaged transcriptomes of neuronal types
- Neural differentiation time course
- Muscular differentiation time course
- Spermatogenesis cell types
- Reprogramming of fibroblasts to iPSCs
- Hematopoietic precursors and cell types