RnoEX6058478 @ rn6
Exon Skipping
Gene
ENSRNOG00000024008 | Cdc25c
Description
cell division cycle 25C [Source:RGD Symbol;Acc:1311875]
Coordinates
chr18:27536255-27541532:-
Coord C1 exon
chr18:27541434-27541532
Coord A exon
chr18:27540212-27540361
Coord C2 exon
chr18:27536255-27536356
Length
150 bp
Sequences
Splice sites
3' ss Seq
CTTGTCAGTATACTTGACAGGAA
3' ss Score
2.16
5' ss Seq
AAGGTTTAG
5' ss Score
2.78
Exon sequences
Seq C1 exon
ATAAACACCAGTTTTAAAACATTGGAATGGGAGGCACCTAGGACCCCGAGGCTTCAAAAAAAGCCTGGCGATCCTCTGGCTTCTCCTCTCTGTGAGCTG
Seq A exon
GAAAATGGAATCTTGCTGGAAAGTGAAATGAAGAATCTGGGCAGTCCCATTACTACAGTTCCAAAACTGAGTAAAAATCTCAAGCTAGAAGACCAAGAAGGGATTTCAGAAGACCCAATGGAGTGTTCCCTGGAAGACCAAGATGCCAAG
Seq C2 exon
GGCTTAAGTCTGAAGAAGACGGTCCCTCTCTGTGGCATGGATGCCATTGAGTTGGAGGAGGAAGATTCTGACTCAGAGCATCTGACTGGTGATTTCTCAAAG
VastDB Features
Vast-tools module Information
Secondary ID
ENSRNOG00000024008-'8-11,'8-10,9-11=AN
Average complexity
A_S
Mappability confidence:
100%=100=100%
Protein Impact
Alternative protein isoforms (Ref)
No structure available
Features
Disorder rate (Iupred):
C1=0.788 A=0.570 C2=0.353
Domain overlap (PFAM):
C1:
PF066178=M-inducer_phosp=FE(18.5=100)
A:
PF066178=M-inducer_phosp=FE(28.3=100)
C2:
PF066178=M-inducer_phosp=PD(11.0=55.9)
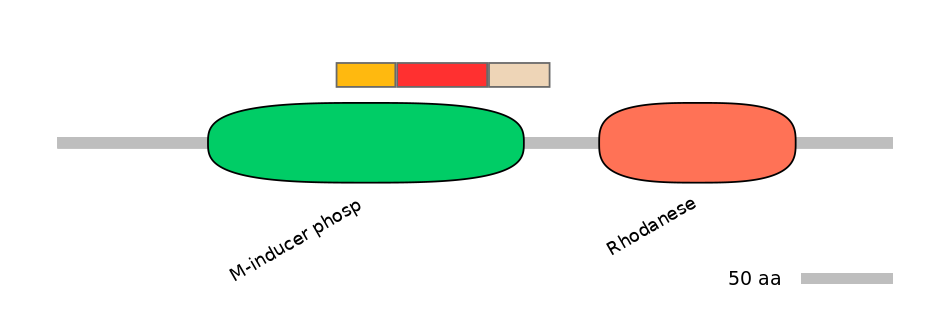
Main Skipping Isoform:
NA
Other Inclusion Isoforms:
NA
Other Skipping Isoforms:
NA
Associated events
Conservation
Chicken
(galGal4)
No conservation detected
Chicken
(galGal3)
No conservation detected
Zebrafish
(danRer10)
No conservation detected
Fruitfly
(dm6)
No conservation detected
Primers PCR
Suggestions for RT-PCR validation
F:
CACCAGTTTTAAAACATTGGAATGGG
R:
TTGAGAAATCACCAGTCAGATGCT
Band lengths:
194-344
Functional annotations
There are 5 annotated functions for this event
PMID: 11842186
Partially encodes an experimentally validated Eukaryotic Linear Motif (ELM). Method: competition binding, mutation analysis. ELM ID: ELMI001012; ELM sequence: NKEND; Overlap: PARTIAL_LEFT
PMID: 11897663
Encodes an experimentally validated Eukaryotic Linear Motif (ELM). Method: alanine scanning, inhibitor. ELM ID: ELMI002160; ELM sequence: EISDELMEFSLKDQE; Overlap: FULL
PMID: 11897663
Encodes an experimentally validated Eukaryotic Linear Motif (ELM). Method: alanine scanning, protein kinase assay. ELM ID: ELMI003363; ELM sequence: MEFSLKD; Overlap: FULL
PMID: 12738781
Encodes an experimentally validated Eukaryotic Linear Motif (ELM). Method: alanine scanning, electrophoretic mobility shift assay, protein kinase assay, western blot. ELM ID: ELMI003363; ELM sequence: MEFSLKD; Overlap: FULL
PMID: 14968113
Encodes an experimentally validated Eukaryotic Linear Motif (ELM). Method: alanine scanning, coimmunoprecipitation, mutation analysis, protein kinase assay. ELM ID: ELMI003380; ELM sequence: EEISDEL; Overlap: FULL
GENOMIC CONTEXT[edit]
INCLUSION PATTERN[edit]