GgaEX0018082 @ galGal4
Exon Skipping
Gene
ENSGALG00000012044 | chAnk2
Description
Uncharacterized protein [Source:UniProtKB/TrEMBL;Acc:F1NG08]
Coordinates
chr4:56338574-56342599:-
Coord C1 exon
chr4:56342335-56342599
Coord A exon
chr4:56340350-56340433
Coord C2 exon
chr4:56338574-56338738
Length
84 bp
Sequences
Splice sites
3' ss Seq
GTCTGTTCTTTGTCATACAGGCC
3' ss Score
8.74
5' ss Seq
AAGGTATTA
5' ss Score
7.04
Exon sequences
Seq C1 exon
TGTCACAACTCCAGGAACAGAGCAGAGGGCAGCAAGCGATACATCTGCACGCTTCAGTGCAACAAAGGAAGAGAGAGAAAAAACATCCCCACAGTCCCCATCATCTGCTCAGAGAGGTGGTTCTCCCATCATCCAAGAGCCTGAAGAGCTCCATCTCCATCAGGATGATCCGTCTCCAAGGAGAACTAGTCTAGTGATAGTGGAGTCAATTGATGAACAGCCAGAAAAGCTTGGAAGCGGCTATGAGGAGGAATCTTTAGAAAAG
Seq A exon
GCCGATAGTATGCCTGAAATGCCACCAGAAACAGTCACAGAAGAACAATACACAGATGAACATGGCCACACAGTGGTGAAGAAG
Seq C2 exon
GTCACTAGAAAGATCATCCGGCGATACGTTTCTCCAGATGGCACTGAAAAAGAAGATATCATTATGCAAGGAACACCACAAAAACCTGTCACAGTTGAAGAAGGAGACGGCTATTCCAAAGTTGTAAAACGAGTCGTGCTGAAGAGTGACAGTGAACAGTCAGAG
VastDB Features
Vast-tools module Information
Secondary ID
ENSGALG00000012044_MULTIEX2-2/2=C1-C2
Average complexity
C1
Mappability confidence:
100%=100=100%
Protein Impact
Alternative protein isoforms (Ref)
No structure available
Features
Disorder rate (Iupred):
C1=1.000 A=0.933 C2=0.834
Domain overlap (PFAM):
C1:
NO
A:
NO
C2:
NO
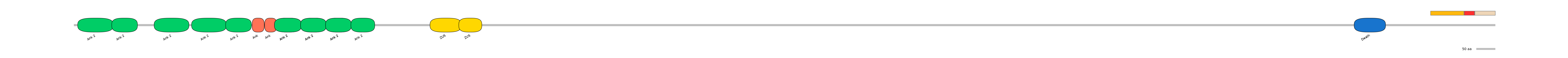
Main Skipping Isoform:
NA
Other Inclusion Isoforms:
NA
Other Skipping Isoforms:
NA
Associated events
Other assemblies
Conservation
Fruitfly
(dm6)
No conservation detected
Primers PCR
Suggestions for RT-PCR validation
F:
AGCGGCTATGAGGAGGAATCT
R:
GTCACTCTTCAGCACGACTCG
Band lengths:
180-264
Functional annotations
There are 0 annotated functions for this event
GENOMIC CONTEXT[edit]
INCLUSION PATTERN[edit]