GgaEX6019266 @ galGal3
Exon Skipping
Gene
ENSGALG00000011007 | F1NN81_CHICK
Description
NA
Coordinates
chr2:31593020-31596740:-
Coord C1 exon
chr2:31596633-31596740
Coord A exon
chr2:31593764-31593898
Coord C2 exon
chr2:31593020-31593113
Length
135 bp
Sequences
Splice sites
3' ss Seq
TTTGTATTTCTTTTAAACAGGAA
3' ss Score
9.81
5' ss Seq
GAGGTCTGA
5' ss Score
2.68
Exon sequences
Seq C1 exon
GCAATAGCACAAAACACAGATCTAAAAGACCGGTTACGCAGAATACATGCCGAATCTATGATCCTTGATTCAGCATCTAATGCAAAATCAAGTCCAGATTTGGTGAAG
Seq A exon
GAAAATTCAAGAGAAGAAAACAGAGGTCTGGTTCACCAGATCTCCAATGAAAGCAGACTGTCTATTGCTGACTCTCTCACAGAATTTTTTGATGCACAGGAAGTCCTACTGTCTGCTAGCTCTTCAGAAAATGAG
Seq C2 exon
GTATCAGATGATGACTCATACGTAAGTGATATCAGTGATAATGTCTCCGAAGATAACCTAAGCAATGACATAGAGAATGAAAGACAAACTTTAG
VastDB Features
Vast-tools module Information
Secondary ID
ENSGALG00000011007-'15-23,'15-22,16-23=AN
Average complexity
A_C2
Mappability confidence:
100%=100=100%
Protein Impact
Alternative protein isoforms (Ref)
No structure available
Features
Disorder rate (Iupred):
C1=0.354 A=0.367 C2=0.797
Domain overlap (PFAM):
C1:
NO
A:
NO
C2:
NO
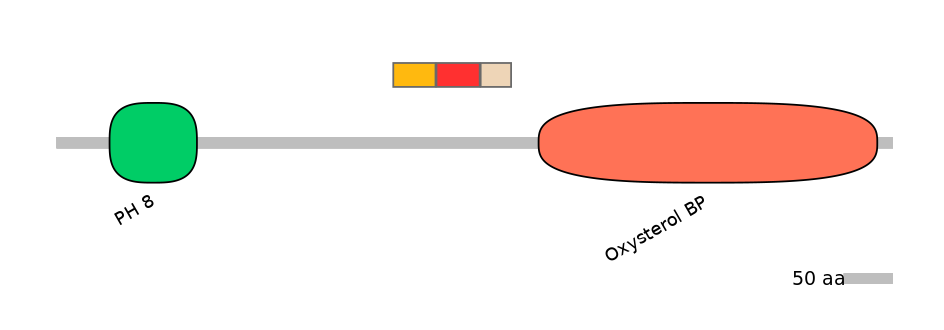
Main Skipping Isoform:
NA
Other Skipping Isoforms:
NA
Associated events
Other assemblies
Conservation
Fruitfly
(dm6)
No conservation detected
Primers PCR
Suggestions for RT-PCR validation
F:
GCAATAGCACAAAACACAGATCT
R:
TGTCTTTCATTCTCTATGTCATTGCT
Band lengths:
194-329
Functional annotations
There are 1 annotated functions for this event
PMID: 17804640
Encodes an experimentally validated Eukaryotic Linear Motif (ELM). Method: classical fluorescence spectroscopy, confocal microscopy, cross linking study, detection by mass spectrometry, green fluorescent protein tag. ELM ID: ELMI001270; ELM sequence: DSLSEFFDAQE; Overlap: FULL
GENOMIC CONTEXT[edit]
INCLUSION PATTERN[edit]