MmuEX6102117 @ mm10
Exon Skipping
Gene
ENSMUSG00000041911 | Dlx1
Description
distal-less homeobox 1 [Source:MGI Symbol;Acc:MGI:94901]
Coordinates
chr2:71528113-71533980:+
Coord C1 exon
chr2:71528113-71530306
Coord A exon
chr2:71530957-71531156
Coord C2 exon
chr2:71532264-71533980
Length
200 bp
Sequences
Splice sites
3' ss Seq
TGCTTCTGTTGTCCCTCCAGGGG
3' ss Score
11.58
5' ss Seq
CAGGTACCT
5' ss Score
8.16
Exon sequences
Seq C1 exon
CGAGTTTGGGGAGCTCGGCAGCATCATGCTTAGACTTTTCAAAGAGACAAACTCCATTTTCTTATGAATGGAAAGTGAAAACCCCTGTTCCGCTTAAATTGGGTTCCTTCCTGTCCTGAGAAACATAGAGACCCCCAAAAGGGAAGCAGAGGAGAGAAAGACCCACACCCAGACCCCGCGAGAAGAGATGACCATGACCACCATGCCAGAAAGTCTCAACAGCCCCGTGTCGGGCAAGGCTGTGTTTATGGAGTTTGGGCCGCCCAACCAGCAAATGTCTCCTTCTCCCATGTCCCACGGGCACTACTCCATGCACTGTTTACACTCGGCGGGCCATTCGCAGCCCGACGGTGCCTACAGTTCGGCCTCATCCTTCTCCCGACCGCTGGGCTACCCCTACGTCAACTCGGTCAGCAGCCACGCGTCCAGCCCCTACATCAGTTCCGTGCAGTCCTACCCCGGCAGCGCCAGCCTCGCCCAGAGCCGCCTGGAGGACCCAG
Seq A exon
GGGCGGACTCCGAGAAGAGTACGGTGGTGGAAGGCGGCGAGGTGCGCTTTAACGGCAAAGGGAAAAAGATCCGTAAACCCAGGACAATTTATTCCAGTTTGCAGTTGCAGGCTTTGAACCGGAGGTTCCAACAAACTCAGTACTTAGCTCTGCCTGAGAGGGCGGAGCTGGCGGCCTCCTTGGGACTCACACAGACGCAG
Seq C2 exon
GTCAAGATATGGTTCCAGAACAAACGCTCCAAGTTCAAGAAGCTGATGAAGCAAGGCGGGGCAGCTCTGGAGGGCAGCGCGCTGGCGAACGGCAGAGCCTTGTCTGCCGGCTCCCCACCGGTACCACCCGGCTGGAATCCGAACTCCTCATCTGGGAAGGGCTCCGGCAGCAGCGCTGGCTCCTACGTCCCCAGCTATACATCATGGTATCCCTCAGCGCACCAAGAAGCTATGCAGCAGCCTCAACTGATGTGAGGTTATCAGTTCGCACTCACCCGCTTCCTGCCCGTCTCCCTTGGCCCGGCCCCTCCCGCCTCCAGGTTCATCCACCTTTTCGCACGCCCTGGGCCAGGAGGAGAAGAGCCCACGAGCGGGAGCAGGGGTGGGGGTGGGGAGCTAAAAATAAAAGAAAAAGGAACAGTGAAGGCAGAGGGCTTGCTGCTTCTCGGAGCCCCTAAAAGTCTGACTCAGCGACCTTTCAGCCCGAATCTGGTTTGGGC
VastDB Features
Vast-tools module Information
Secondary ID
ENSMUSG00000041911-'7-10,'7-8,9-10
Average complexity
S
Mappability confidence:
100%=100=100%
Protein Impact
ORF disruption upon sequence exclusion
No structure available
Features
Disorder rate (Iupred):
C1=0.548 A=0.348 C2=0.512
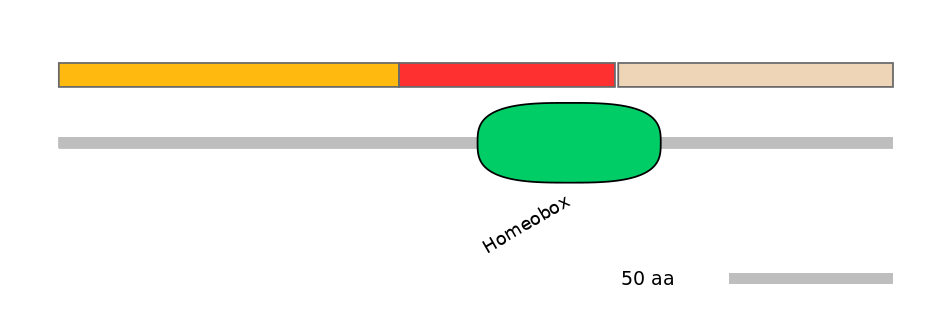
Domain overlap (PFAM):
C1:
NO
A:
PF0004624=Homeobox=PU(73.7=62.7)
C2:
PF0004624=Homeobox=PD(22.8=15.3)
Main Skipping Isoform:
NA
Other Skipping Isoforms:
NA
Associated events
Other assemblies
Conservation
Primers PCR
Suggestions for RT-PCR validation
F:
GGGCAAGGCTGTGTTTATGGA
R:
GGAGCGTTTGTTCTGGAACCA
Band lengths:
299-499
Functional annotations
There are 0 annotated functions for this event
GENOMIC CONTEXT[edit]
INCLUSION PATTERN[edit]
SPECIAL DATASETS
- Ribosome-engaged transcriptomes of neuronal types
- Neural differentiation time course
- Muscular differentiation time course
- Spermatogenesis cell types
- Reprogramming of fibroblasts to iPSCs
- Hematopoietic precursors and cell types