GgaEX0027170 @ galGal4
Exon Skipping
Gene
ENSGALG00000002198 | FHOD1
Description
FH1/FH2 domain-containing protein 1 [Source:RefSeq peptide;Acc:NP_001012792]
Coordinates
chr11:1336785-1350244:+
Coord C1 exon
chr11:1336785-1337084
Coord A exon
chr11:1347893-1348045
Coord C2 exon
chr11:1350088-1350244
Length
153 bp
Sequences
Splice sites
3' ss Seq
GCTCCCTGTCCTCTTTGCAGAAA
3' ss Score
10.76
5' ss Seq
GAGGTGAGA
5' ss Score
7.66
Exon sequences
Seq C1 exon
CCCGCTTACACCCGCGGAGGGCGCCCGAGCGCGGGGCGGTGCCGGCTCAGAGCCGGGGCGGTGCCGGGGGCTGTCCCGATGCCACCCCCGGAGCTCCGGGCGCGGTGCGGCGGCGGTGGCGGTGCCGCCATGGCGGAGGCTGCGGTCCCGTGCCGGGTGCAGTACCTGGAAGACTCCGATCCCTTCGGCTGCGGCAGTTTCCCGGAGCCCCGACGGGCGCCTGTTTACGCCGTGGAGGAAGCGTTGGCCCTGGGAGCGCAGCTGCCCGCGGTGCACCGGCTGCTGGGAGCGCCGCTGCCG
Seq A exon
AAAGTTACTGATGGAAAGAAAGTGGTGGTGGTGCTGGATCCCAAGAGGAGCAACGCCATCAACATTGGCCTCACGGTGCTGCCACCTGTGCACATCATCAAGACAGCCGTGCTCAACTTTGATGAGTTTGCAGTCAGTAAGGAAGGGATTGAG
Seq C2 exon
ACCGAGAAGTTCTCTGGCATCACCGAGGCATCACCACCCGCCGTGGTGTCAGCCAGCCCCACGCAGCAGGAGCAGGCAGAGGTGGGCCACGAGAGCATGAAGAGCGTCCTGAGCTCGCCCACGGATGTCCCTGCCCGCCGCAGTCGGGGTGGCCGCG
VastDB Features
Vast-tools module Information
Secondary ID
ENSGALG00000002198_MULTIEX1-13/18=C1-C2
Average complexity
C3
Mappability confidence:
100%=100=100%
Protein Impact
Alternative protein isoforms (Ref)
No structure available
Features
Disorder rate (Iupred):
C1=0.162 A=0.000 C2=0.962
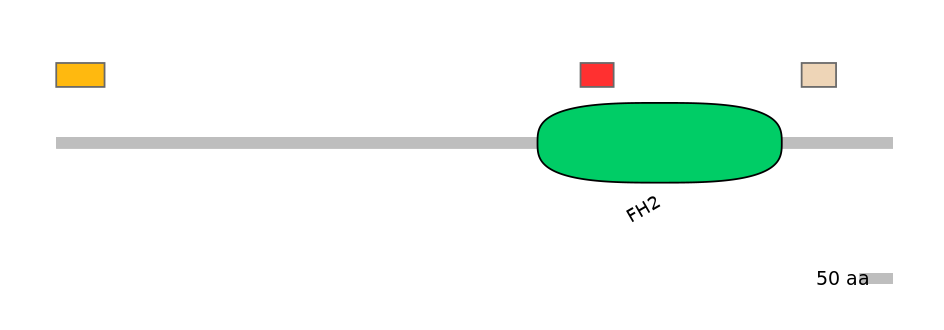
Domain overlap (PFAM):
C1:
NO
A:
PF0218118=FH2=FE(13.5=100)
C2:
NO
Main Skipping Isoform:
NA
Other Inclusion Isoforms:
NA
Other Skipping Isoforms:
NA
Associated events
Other assemblies
Conservation
Fruitfly
(dm6)
No conservation detected
Primers PCR
Suggestions for RT-PCR validation
F:
CCTGGAAGACTCCGATCCCTT
R:
CAGGACGCTCTTCATGCTCTC
Band lengths:
247-400
Functional annotations
There are 0 annotated functions for this event
GENOMIC CONTEXT[edit]
INCLUSION PATTERN[edit]