HsaEX6063938 @ hg38
Exon Skipping
Gene
ENSG00000107147 | KCNT1
Description
potassium sodium-activated channel subfamily T member 1 [Source:HGNC Symbol;Acc:HGNC:18865]
Coordinates
chr9:135769947-135771095:+
Coord C1 exon
chr9:135769947-135770055
Coord A exon
chr9:135770298-135770447
Coord C2 exon
chr9:135770857-135771095
Length
150 bp
Sequences
Splice sites
3' ss Seq
GGCCCCGCCCTGGCCCACAGGGA
3' ss Score
7.64
5' ss Seq
GAAGTAAGG
5' ss Score
6.38
Exon sequences
Seq C1 exon
ACCACGTGGTGTGTGAGGAGGAGTGCAAGTACGCCATGCTGGCGCTGAACTGCATCTGCCCGGCGACCTCCACCCTCATCACCCTGCTGGTGCACACGTCCCGCGGCCA
Seq A exon
GGAGGGACAGGAGTCTCCGGAGCAGTGGCAGCGCATGTATGGGCGCTGCTCCGGCAACGAGGTGTACCACATCCGCATGGGTGACAGCAAGTTCTTCCGCGAGTACGAGGGCAAGAGCTTCACCTACGCGGCCTTCCACGCCCACAAGAA
Seq C2 exon
GTATGGCGTGTGCCTCATCGGGCTGAAGCGGGAGGACAACAAGAGCATCCTGCTGAACCCGGGGCCCCGGCACATCCTGGCCGCCTCTGACACCTGCTTCTACATCAACATCACCAAGGAGGAGAACTCGGCCTTCATCTTCAAGCAGGAGGAGAAGCGGAAGAAGAGGGCCTTCTCGGGGCAGGGGCTGCACGAGGGTCCGGCCCGCCTGCCCGTGCACAGCATCATCGCCTCCATGG
VastDB Features
Vast-tools module Information
Secondary ID
ENSG00000107147_MULTIEX1-2/2=1-C2
Average complexity
S
Mappability confidence:
100%=100=100%
Protein Impact
Alternative protein isoforms (Ref)
No structure available
Features
Disorder rate (Iupred):
C1=0.000 A=0.098 C2=0.108
Domain overlap (PFAM):
C1:
PF0349313=BK_channel_a=FE(34.0=100)
A:
PF0349313=BK_channel_a=FE(47.2=100)
C2:
PF0349313=BK_channel_a=PD(8.5=11.1)
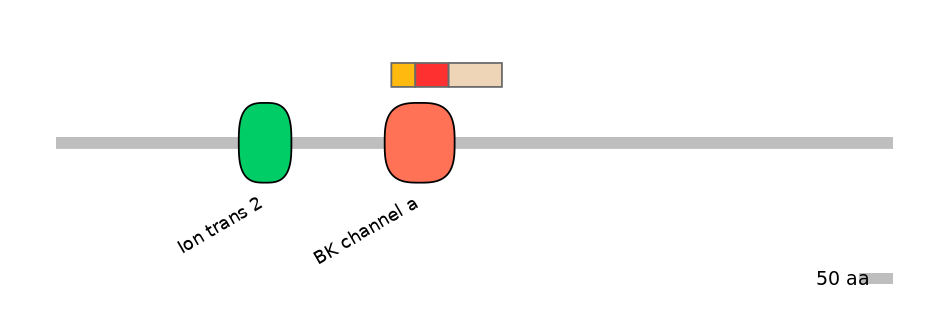
Main Skipping Isoform:
NA
Other Inclusion Isoforms:
Other Skipping Isoforms:
NA
Associated events
Other assemblies
Conservation
Primers PCR
Suggestions for RT-PCR validation
F:
GAGGAGTGCAAGTACGCCATG
R:
CGCTTCTCCTCCTGCTTGAAG
Band lengths:
251-401
Functional annotations
There are 0 annotated functions for this event
GENOMIC CONTEXT[edit]
INCLUSION PATTERN[edit]
SPECIAL DATASETS
- Autistic and control brains